Summary of the Overall Objective of the Study
Dishonesty in the food sector is a rampant global problem. The most controversial case of food fraud was the European ‘horsemeat scandal’ that took place in 2013 (FSAI 2013). The dishonest addition of extra meat products to food without acknowledging their presence has some negative implications for consumers, including financial, ethical, and religious consequences. For instance, the unscrupulous blending of food items with horse meat or pork may contravene the religious beliefs of individuals who are against the consumption of such meats.
Consumers are aware of such possibilities, which necessitate the availability of reliable means of checking food for fraudulent blending with undeclared meat products. The commonly used techniques such as enzyme-linked immunosorbent assay (ELISA) and polymerase chain reaction (PCR) have known shortcomings such as reduced sensitivity or specificity during the analysis of processed food items.
Therefore, the main purpose of the study was to create an optimized technique for the identification of contaminated pork and horse in various food items using multiple reaction monitoring (MRM) recognition of species-specific tryptic marker peptides (Von Bargen, Brockmeyer & Humpf 2014).
The Rationale for the Use of Proteomic Techniques
Currently, species certification uses ELISA and PCR techniques. ELISA may serve as a qualitative or quantitative method during food production. This method can also be applied to processed samples and has been reported to have a limit of detection of 0.5% (Von Bargen, Brockmeyer & Humpf 2014). However, the specificity of ELISA in samples that have undergone extensive processing is questionable, which leads to false positive or false negative upshots. Moreover, multiplexing, which is the simultaneous identification and measurement of more than one species in a single experiment, cannot be done effectively using ELISA.
PCR is an alternative method for regular meat speciation in laboratories. Contemporary reports involving PCR indicate that the method has exceptional sensitivity in processed foods (Ali et al. 2015; Hou et al. 2015; Karabasanavar et al. 2014). However, the main limitation of PCR is that DNA is highly sensitive to food processing. As much as using mitochondrial DNA templates can augment the effectiveness of PCR in processed commodities, it has also been reported that the DNA template can be broken down by slight heating (Ali et al. 2015; Karabasanavar et al. 2014). Nonetheless, PCR permits the development of multiplex techniques, thereby making it advantageous over ELISA.
In recent times, proteomics techniques have been used in meat certification to circumvent the problems associated with ELISA and PCR. Examples include mass spectrometry (MS)-based proteomics techniques where precise proteins are identified using MS techniques. The proteins are produced following tryptic digests of protein samples. The basis of proteomics in species identification is the stability of the primary structure of proteins against processing.
Besides, some degree of protein disintegration does not have a substantial impact on the identification of species-specific peptides as opposed to the shearing of DNA for PCR detection (Watson et al. 2015; Naveena et al. 2018). Also, short peptide sequences are less likely to undergo fragmentation compared to the degradation of long DNA templates. MS instrumentation has undergone various enhancements that improve its application in targeted proteomics methods as well as the finding of species-specific peptides through MRM.
Summary of Proteomic Methodology and Results
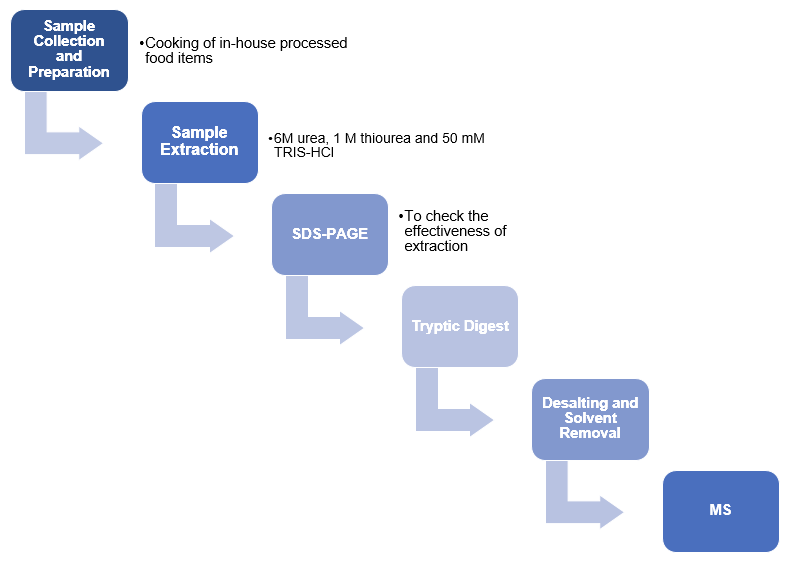
Meat samples were purchased and stored at -20oC until subsequent analysis. In-house processed samples were prepared by slicing into thin pieces and mincing using an electric grinder. The ground meat was made into meatloaves, frikadeller, and meatballs by cooking using respective recipes until core temperatures of 100oC were attained. The food items were pre-crushed in freezing bags, frozen in nitrogen, and pulverized for 5 minutes. Sample mixtures were prepared by contaminating ground beef with ground pork or horse meat. On the other hand, commercially processed samples were cut into small pieces, washed to get rid of pasta sauce, and pulverized.
Protein extraction was done using 1 g of each sample in 10 ml of extraction buffer comprising “6 M urea, 1 M thiourea and 50 mM TRIS-HCl” regulated to pH 8 (Von Bargen, Brockmeyer & Humpf 2014, p. 9429). All samples were vortexed for 20 seconds. However, those with large concentrations of connective tissue were dissolved for a total of 110 seconds at 9500 to 20 500 rpm. The extracted samples were then centrifuged for 1 hour at 4 °C at 12 000g followed by the collection of a 100-μL aliquot of the supernatant in Eppendorf tubes.
Precast sodium dodecyl sulfate (SDS) polyacrylamide gradient gels (4 to 20%) were run to determine the efficiency of the extraction. The extracts were diluted using water and sample buffer followed by denaturation at 95 °C for 300 seconds. Thereafter, the samples were applied to the SDS-gels that were focused for 20 minutes at 100 V before running at 140 V. Coomassie brilliant blue dye was used to stain the gels for 25 minutes, followed by visualization and imaging. A standard protocol was used to perform tryptic digest, which was followed by desalting and solvent removal. The samples were then re-dissolved in ACN/H2O and analyzed by MS.
It was noted that heat treatment, for instance, frying and boiling lowered the solubility of target proteins. However, the area, thiourea, and Tris-HCl combination yielded the best outcomes. Maillard reaction in fried meat products produced a dark brown color, which was reflected in the resultant peptide solutions as a yellow to brown color suggesting changes in the protein matrix. Therefore, lower sensitivities were noted for harshly processed samples compared to mildly processed ones (boiling).
Out of the 23 samples that were analyzed, only 1 did not meet the expected standards. Horse-specific peptide markers using MRM experiments were detected in a corned beef sample shown to contain only beef material. Droplet digital PCR was used to verify the results. Overall, the technique facilitated the identification of 0.24% of horse or pork in a meat medium.
The Potential Usefulness or Application of Research Findings
The research findings show that the modified technique was sensitive in the identification of trace amounts of horse meat. This method circumvented the previously mentioned limits of detection in ELISA and PCR. The technique could be implemented without the need for special proteomics infrastructure, for example, off-gel fractionation of 2D-PAGE. Consequently, the protocol is feasible for laboratories without specific proteomics expertise. Therefore, this technique can be used in food industries and quality control organizations for meat authentication.
Subsequent Publications that Have Cited or Advanced this Work
Current statistics on Google Scholar indicate that this article has been cited 77 times since it was published. These papers mainly focus on the authentication of processed meat. For example, Balog et al. (2016) use rapid evaporative ionization mass spectrometry to ascertain the species of origin of meat products. Montowska et al. (2015) use this work as a basis for the development of their work where they identified heat-labile peptide markers in meat using ambient liquid extraction surface analysis mass spectrometry (LESA-MS). Furthermore, three pork and horse markers used in this paper are the same as those applied by Von Bargen, Brockmeyer, and Humpf (2014).
Conclusion
Proteomics has proven to be a reliable means of identifying contaminating meat species in food samples. Its strengths are based on the ability to circumvent the destruction of DNA during food processing, high sensitivity, and limits of detection. Future studies should focus on improving this technique.
Reference List
Ali, ME, Razzak, MA, Hamid, SBA, Rahman, MM, Al Amin, M & Rashid, NRA 2015, ‘Multiplex PCR assay for the detection of five meat species forbidden in Islamic foods’, Food Chemistry, vol. 177, pp. 214-224.
Balog, J, Perenyi, D, Guallar-Hoyas, C, Egri, A, Pringle, SD, Stead, S, Chevallier, OP, Elliott, CT & Takats, Z 2016, ‘Identification of the species of origin for meat products by rapid evaporative ionization mass spectrometry’, Journal of Agricultural and Food Chemistry, vol. 64, no. 23, pp. 4793-4800.
FSAI 2013, FSAI survey finds horse DNA in some beef burger products. Web.
Hou, B, Meng, X, Zhang, L, Guo, J, Li, S & Jin, H 2015, ‘Development of a sensitive and specific multiplex PCR method for the simultaneous detection of chicken, duck and goose DNA in meat products’, Meat Science, vol. 101, pp. 90-94.
Karabasanavar, NS, Singh, SP, Kumar, D & Shebannavar, SN 2014, ‘Detection of pork adulteration by highly-specific PCR assay of mitochondrial D-loop’, Food Chemistry, vol. 145, pp. 530-534.
Montowska, M, Alexander, MR, Tucker, GA & Barrett, DA 2015, ‘Authentication of processed meat products by peptidomic analysis using rapid ambient mass spectrometry’, Food Chemistry, vol. 187, pp. 297-304.
Naveena, BM, Jagadeesh, DS, Kamuni, V, Muthukumar, M, Kulkarni, VV, Kiran, M & Rapole, S 2018, ‘In‐gel and OFFGEL‐based proteomic approach for authentication of meat species from minced meat and meat products’, Journal of the Science of Food and Agriculture, vol. 98, no. 3, pp. 1188-1196.
Von Bargen, C, Brockmeyer, J & Humpf, HU 2014, ‘Meat authentication: a new HPLC–MS/MS based method for the fast and sensitive detection of horse and pork in highly processed food’, Journal of Agricultural and Food Chemistry, vol. 62, no. 39, pp. 9428-9435.
Watson, AD, Gunning, Y, Rigby, NM, Philo, M & Kemsley, EK 2015, ‘Meat authentication via multiple reaction monitoring mass spectrometry of myoglobin peptides’, Analytical Chemistry, vol. 87, no. 20, pp. 10315-10322.