Introduction
Antibiotics are antimicrobial agents that aid in the treatment or deterrence of bacterial infections (1). These chemical agents are sometimes referred to as “antibacterial.” Their mode of action entails either killing the bacteria or inhibiting their growth. Antibiotics that prevent the growth of bacteria are called bacteriostatic antibiotics, while those that kill bacteria are known as bactericidal antibiotics (1). Bacteriostatic antibiotics interfere with the essential physiological processes of bacterial cells including the production of proteins, as well as the replication of DNA and RNA. By doing so, the bacteria are unable to multiply and carry out their metabolic activities, permitting the body’s immune system to clear the infection. Bactericidal antibiotics, on the other hand, kill bacteria by destroying the cell membrane or cell wall, thus allowing the cell contents to leak out and leading to the death of the bacterial cells.
Disinfectants are described as antimicrobial agents that are applied to non-living objects to get rid of any microorganisms present on the objects. The mode of action of disinfectants involves interfering with the metabolism of bacteria and the integrity of the bacterial cell walls. However, disinfectants are not one hundred percent effective as they are unable to eliminate resistant spores.
Antimicrobial susceptibility testing is a process of finding out which specific antibiotics a given bacterial species is sensitive to. Different bacterial species demonstrate different sensitivities to various antibiotics. For example, the emergence of methicillin-resistant Staphylococcus aureus is a global problem in clinical medicine due to the bacterial species’ resistance to beta-lactam antibiotics and penicillin (2). Therefore, susceptibility testing is beneficial in determining the most effective antibiotic to manage a bacterial infection in a living cell. This method can also be used to identify certain bacteria. One of the commonly used techniques for determining antimicrobial sensitivity is the disc diffusion method.
The benefits of this approach are that it is convenient, effective, and economical. To complete this test, a growth medium is poured evenly on a plate alongside the bacterial isolate of interest, which has been watered down to a standard concentration. Commercially made discs pre-infused with a known strength of a specific antibiotic are then spread evenly and pressed lightly on the surface of the agar. The antibiotic diffuses outward from the discs, creating a gradient of antibiotic concentration in the agar such that the highest concentration is located near the disc, while lower concentrations are found further away from the disc. The bacterial growth around each disc is then observed following incubation. Susceptible isolates show a clear zone devoid of growth around a given disc.
The purpose of this experiment is to determine the susceptibility of three bacteria to different antibiotics and disinfectants. It was hypothesized that S. aureus and Pseudomonas aureginosa would exhibit similar patterns of antibiotic and disinfectant susceptibility.
Materials & Methods
Different antibiotic solutions including ampicillin (AM10), chloramphenicol (C30), kanamycin (K30), nalidixic acid (NA30), novobiocin (NB5), gentamicin (GM10), tetracycline (TE30), penicillin (P10), sulfamethazole/trimethoprim (SXT), CID5, CF30, and S10 were prepared. Bacterial culture tubes containing three bacterial strains—Escherichia coli, Staphylococcus aureus, and Pseudomonas aureginosa—were flicked to mix the contents. Cotton swabs were aseptically dipped one time in the bacterial broth, after which the swabs were spread uniformly on three Petri dishes containing agar.
Twelve discs were aseptically transferred to the 12-well tray by placing one disc per well. One drop of disinfectant or antibiotic was added to each disc. The impregnated discs were then placed on the plates following bacterial plating of the bacteria in such a way that each disc centered one quadrant per plate of the three square plates. The plates were then inverted and incubated at 37° C for three days. On the third day, the diameters of the zones of inhibition of all antibiotics and disinfectants were measured in millimeters and recorded. The inhibition zones were then compared to a standard interpretation chart and used to classify the bacteria isolates as sensitive, intermediately sensitive, or resistant.
Results
Table 1: The mean zones of inhibition for the three bacterial species growing on different antibiotics.
Key: + sensitive; – resistant; * intermediate.
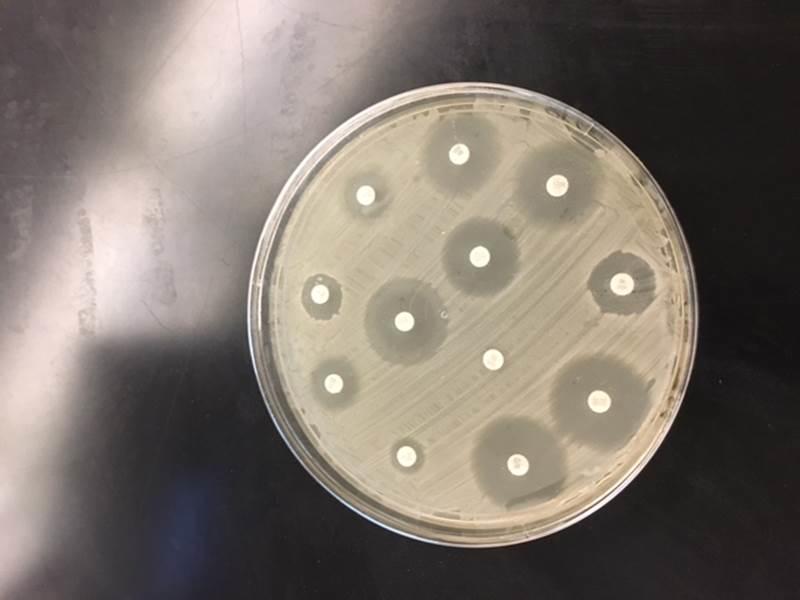
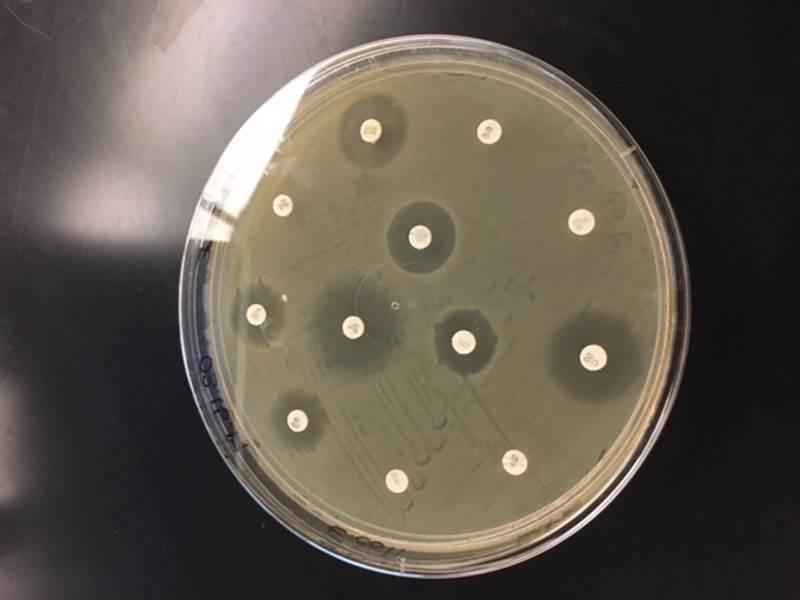
The clear regions surrounding the discs indicate the zones of inhibition, whereas the opaque areas around the discs show that the bacteria are resistant to the antibiotics.
Discussion
E. coli is a Gram-negative, rod-shaped bacterium prevalent in the lower intestines of most endothermic living organisms (1). E. coli is a facultative anaerobe. S. aureus, on the other hand, is a Gram-positive bacterium whose cells have a round shape. S. aureus is a common microflora of the skin, nose, and respiratory system. P. aeruginosa is a Gram-negative bacterium that is responsible for plant and animal disease. Its cells are rod-shaped. All three of these microbes were resistant to penicillin and novobiocin. In contrast, they were all susceptible to gentamicin, kanamycin, tetracycline, and sulfamethoxazole. There was an intermediate sensitivity of S. aureus and P. aureginosa to chloramphenicol. S. aureus was the only microbe that was susceptible to ampicillin and resistant to nalidixic acid. The findings supported the hypothesis that E. coli and P. aureginosa would exhibit similar antibiotic susceptibility patterns given that they are both Gram-negative.
The mechanisms of different antibiotics determined which microbes were susceptible to it. For example, ampicillin irreversibly inhibits the enzyme transpeptidase, whose role is critical to the synthesis of bacterial cell walls. Transpeptidase inhibition interferes with the final step in bacterial cell wall synthesis during binary fission. Consequently, cell lysis occurs, leading to cell death. Chloramphenicol slows down bacterial growth by preventing protein synthesis, particularly the elongation of the protein chain, by blocking the peptidyl transferase action of the bacterial ribosome. Kanamycin and gentamicin are aminoglycosides whose mode of action entails binding the bacterial 30S ribosomal subunit and thwarting the actions of t-RNA. Consequently, protein synthesis is inhibited. Aminoglycosides are useless against anaerobic bacteria, which explains why the two antibiotics were effective against all three microbes.
Nalidixic acid hinders DNA synthesis in Gram-negative bacteria, which accounts for its inhibition of E. coli and P. aureginosa (3). Conversely, novobiocin is an aminocoumarin that impedes the action of DNA gyrase in bacteria, thus impairing the transduction of energy. Tetracyclines inhibit protein synthesis and are effective against most aerobic bacteria. Penicillin exerts its bactericidal actions by hampering the cross-linking of peptidoglycan during cell wall synthesis. Therefore, it is effective against both Gram-positive and Gram-negative bacteria. Sulfamethoxazole/trimethoprim obstruct bacterial production of dihydrofolate, thus affecting the synthesis of proteins and nucleic acids (4), which explains its effectiveness against these three bacteria. The data in this experiment are consistent with the literature, and it was concluded that E. coli and P. aureginosa demonstrated similar antibiotic sensitivity patterns.
References
- Tortora GJ, Funke BR, Case, CL. 2015. Microbiology: An introduction. San Francisco, CA: Pearson Benjamin Cummings.
- Tang SS, Apisarnthanarak A, Hsu LY. 2014. Mechanisms of β-lactam antimicrobial resistance and epidemiology of major community and healthcare-associated multidrug-resistant bacteria. Adv Drug Deliv Rev 78: 3-13. Web.
- Aldred KJ, Kerns RJ, Osheroff N. 2014. Mechanism of quinolone action and resistance. Biochemistry 53: 1565-1574. Web.
- Livermore DM, Mushtaq S, Warner M, Woodford N. 2014. Comparative in vitro activity of sulfametrole/trimethoprim and sulfamethoxazole/trimethoprim and other agents against multiresistant Gram-negative bacteria. J Antimicrob Chemother 69: 1050-1056. Web.