Introduction
This practice was done in two parts. The first part involved the separation of mitochondria from rabbit liver using differential and density gradient centrifugation. The differential and density gradient centrifugation technique involved centrifuging cell homogenate at low speed to get rid of cellular debris and large organelle components. The last step of centrifugation involved centrifuging pellets at high speed to obtain mitochondria. The technique utilized density properties of cell components.
The second part of the practice concentrated on assaying for mitochondrial enzymes and proteins to ascertain that mitochondria were isolated from the rabbit liver. One of the techniques used was based on the enzymatic activity of a mitochondrial enzyme known as succinate dehydrogenase. This enzyme was used as a marker for mitochondria. The other technique used was SDS-PAGE to analyze mitochondrial fractions for the presence of proteins. The protein sample was intact in its four structures (primary, secondary, tertiary and quaternary structures). The purpose of the SDS was to disrupt the secondary and tertiary conformations of the sample proteins. The charge forced the proteins to open up and became straight. Heating was done to make the SDS enter the hidden water-hating areas of the protein sample. Quaternary structures were disrupted by beta-mercaptoethanol, which acted by reducing the chemical bonding within the conformation (it acted as a reducing agent). The SDS-PAGE technique utilized differences in size and intensity of protein bands on a gel.
Aims
The two parts of the experiment used rabbit liver as the sample. The first aim of the experiment was to isolate mitochondria using differential gradient and density centrifugation. The second aim of the experiment was to assay for a specific marker (succinate dehydrogenase) located in mitochondrion to ascertain that mitochondria were isolated from the rabbit liver sample. The enzyme was used as a marker to detect mitochondria in the rabbit liver. The third aim of the experiment was to test for mitochondrial proteins using SDS-PAGE to ascertain that, in fact, mitochondria were isolated from the rabbit liver. The experiment used the aforementioned molecular biology techniques to achieve these aims.
Methods
Isolation of mitochondria from rabbit liver
Five (5) tubes, with lids, were labelled as WCL (whole cell lysate), SN1 (supernatant 1), SN2 (supernatant 2), SN3 (supernatant 3) and MITO (mitochondrial fraction). Liver tissue homogenate was centrifuged to obtain cell lysate and pellet. Cell lysate (100 µL) was stored on ice and the pellet of the cell breakdown was centrifuged at 1,000xg at 4°C for 10 minutes. The supernatant was transferred to the MITO tube and the pellet discarded. The supernatant was centrifuged at 10,000xg for 10 minutes at 4°C. The supernatant was stored on ice in the SN1 tube. The pellet was mixed thoroughly with 1000 µL of mito-buffer. The resulting mixture was centrifuged at 12,000xg at 4°C for 10 minutes. The supernatant obtained was transferred to the SN2 tube and stored on ice. The pellet at the bottom of tube was mixed thoroughly with 1000 µL mito-buffer and centrifuged at 12,000 xg for 10 minutes at 4°C. The supernatant was placed in the SN3 tube. The pellet remaining at the bottom of the tube was mixed thoroughly with 400µL of mito-buffer. This was the mitochondrial pellet that was placed in the MITO tube.
Succinate Dehydrogenase Assay
To each of the tubes (WCL, SN1, SN2, and MITO) containing the cell fractions obtained, 300 µL of Succinate solution was added. Blanks tubes were labelled as the mentioned tubes but with the word BLANK at the end of the labelling. Also, Succinate solution was added to these tubes without samples. The tube contents were mixed well and incubated at body temperature of 37°C for 10 minutes. Stop solution was added to the BLANK tubes before start of incubation while the solution was added to the tubes with samples after incubation process. The contents of the tubes were centrifuged for 2 minutes at maximum speed. The supernatant of the tubes were placed on glass cuvettes and absorbance for each sample measured against its blank using a spectrophotometer. The blank was removed then the sample tube with the corresponding label measured for absorbance. The enzyme activity was compared among the four tubes of samples by graphing absorbance units of the samples, using standards of absorption of each fraction at 490 nm.
Analysis by SDS-PAGE and gel visualizing
From each sample tubes WCL SN1, SN2 and MITO, 60 µg of the samples were transferred to microfuge tubes. Enough water was added to the microfuge tubes to top up the volume to 20µL. 6µL of SDS PAGE buffer was added to the tubes and the contents mixed thoroughly. The tubes were briefly centrifuged to bring all their contents to the bottom to ensure that there were no drops of contents on the lids and walls of the tubes. The samples were incubated at room temperature for 15 minutes before they were loaded onto the gel. Electrophoresis was carried out at 200 volts until dye was seen at the bottom of the gel. The samples loaded onto the gel migrated from one electrode to the other. The samples migrated along the gel depending on their molecular sizes. The heavier proteins moved faster than the lighter proteins along the gel. The electrophoresis was stopped and apparatus dismantled. Gel was activated for viewing by illuminating UV on it then visualized using a special Gel viewer.
Results
Proteins were successfully isolated from the sample fractions and their amounts were estimated using the graph shown in Figure1. The SN1 fraction of the sample had the highest amount of proteins (3802.23µg) followed by the mitochondrial fraction with 744.99µg. The SN3 fraction had the lowest number of proteins estimated to be 164.65µg. The SN2 fraction of the sample had 248.34µg of sample proteins.
The sample fractions were assayed for activity of the enzyme succinate dehydrogenase using a spectrophotometer which gave levels of absorbance as shown in Table1. The fractions WCL, SN1, SN2 and MITO had absorbance values of 0.146, 0.068, 0.065 and 0.155 respectively. The absorbance of the samples was measured at 490 nm. To estimate the molecular weights of the samples, mobility of the marker bands was plotted against their molecular weights in kD (Figure2). From this graph the molecular weights of the bands found in the sample fractions were estimated. The mitochondrial fraction was found to contain a strong band with a molecular weight of 150 kD. The WCL fraction only contained the 250 kD and 150 kD (which was not strong) bands. Table2 shows the bands that were unique for each of the sample fractions as they were observed on the gel photo (Figure4).
Figures and Tables
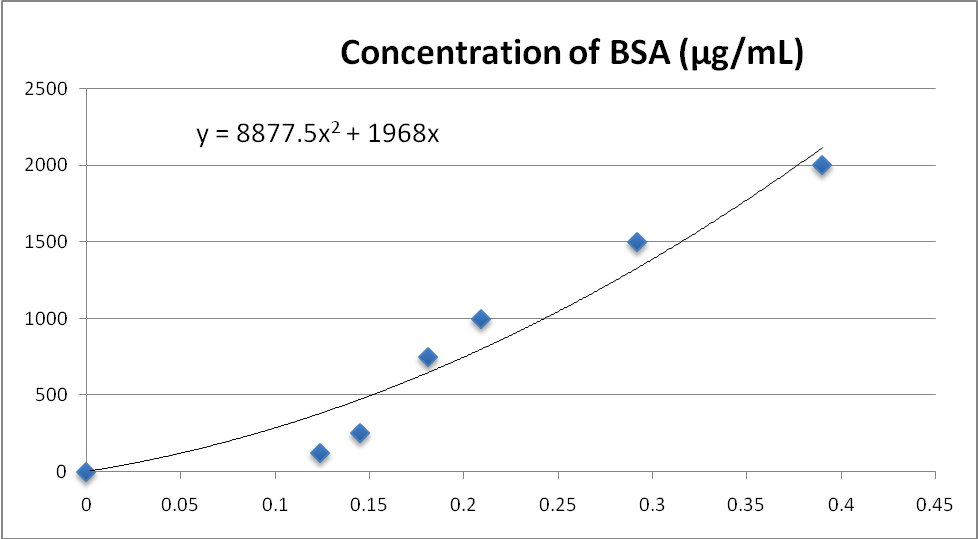
Graph showing the relationship between concentration of BSA (µg/mL) on the Y-axis and absorbance on the X-axis. The graph was used to give best line with the equation y = 8877.5x2 + 1968x which was used to calculate the amounts of proteins extracted from the rabbit liver sample. The graphing was done on Microsoft Excel.
Table 1.
The table was used compare the absorbance values of the four fractions of the sample that were assayed for succinate dehydrogenase. Among the four fractions, the mitochondrial fraction had the highest absorbance while the SN2 had the lowest value. This was an indication that the mitochondrial fraction had the highest concentration of the enzyme succinate dehydrogenase. The WCL fraction was the second fraction with the highest amount of the enzyme succinate dehydrogenase.
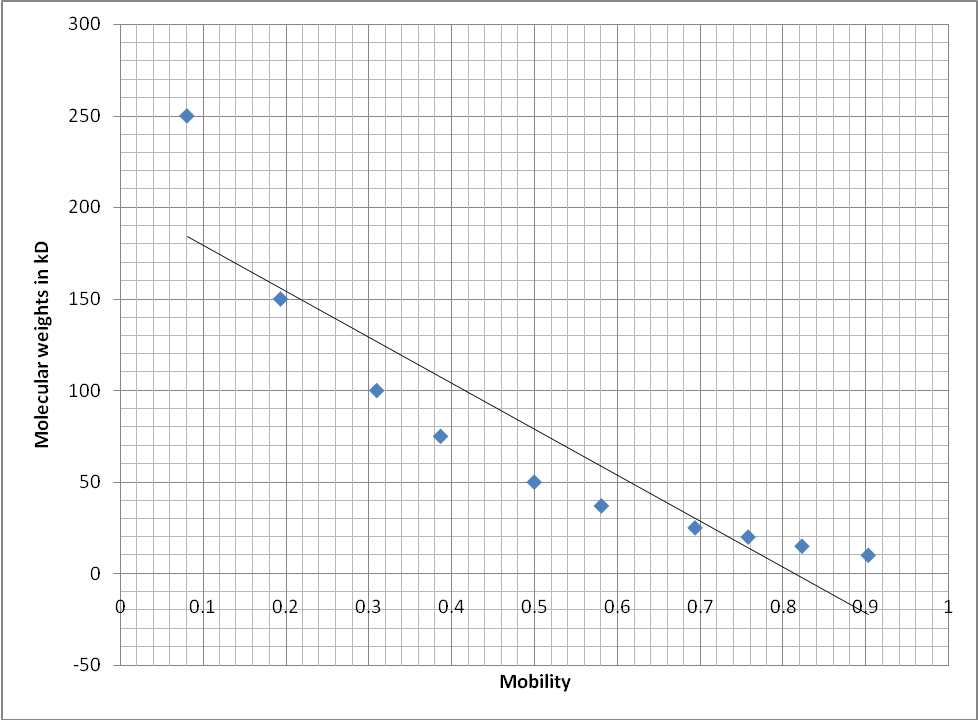
The graph was used to display the relationship between molecular weights of the standards and the mobility of the bands. Mobility was calculated by dividing the distance moved by respective bands by the total length of the gel. This was an inverse relationship indicating that the heavier the band was the shorter the distance it migrated from the origin of the acrylamide gel. On the other hand, the lighter the band the longer the distance it moved from the origin of the gel.
Table 2.
The table shows the distinct bands among the mitochondrial, WCL and SN fractions. The WCL fraction contained only the 249kD and 150kD bands. The 250kD band was common in the SN1 and SN2 fractions of the sample. Among all the fractions, the 150kD band was strongest in the mitochondrial fraction of the sample. However, the 150kD, 100kD and 74kD band was visualised in the SN1, SN2 and MITO fractions of the sample.
Discussion
The results obtained from this experiment were important in the study of cellular and sub-cellular components and their functions. Different parts of the cell have been characterised by unique chemical components used as the markers for the parts. Studies have always identified mitochondria with the enzyme succinate dehydrogenase, a nitric acid cycle enzyme located in the inner membrane of mitochondrion (Speers & Cravatt, 2004, p.536). Mitochondria have been regarded as the organelles in eukaryotic cells that supply the cells with energy (Ferrer, 2009, pp.427-430). Enzymatic pathways have been suggested to be involved in processes that yield energy for the cells (Gray, 2012, p.15).
Contamination might have been encountered in the course of running this experiment. The contamination might have resulted in wrong results which resulted in wrong conclusions regarding the sample results. For example, bands were observed in lanes whose wells were loaded with loading buffers without samples. This might have happened during loading and handling of the sample fractions. Errors in centrifugation might have resulted in poor separation of sample proteins to their respective fractions. This was shown by the various band sizes that were visualised in all the fractions. Other studies identified unique bands in respective sample fractions.
This experiment confirmed results from previous studies which associated succinate dehydrogenase with the mitochondrial fraction of eukaryotic cells. The other studies also used spectrophotometer to assay for the amounts of succinate dehydrogenase in samples. The experiment also confirmed results from previous studies which reported that amounts of proteins extracted in SN3 fraction were less than those extracted in the MITO fraction (Murphy, 2009, p.12). Study reports have indicated that higher amounts of succinate dehydrogenase in SN3 than MITO fractions could be an indication of contamination of SN3 sample fraction with MITO fraction components. Results from this experiment confirmed reports from previous studies which indicated that SDS-PAGE was an efficient system of resolving proteins based on their molecular sizes (Schägger, 1995, p.190). The SDS-PAGE separated proteins with the heavy proteins moving only a short distance form the origin of the gel while the lighter proteins moved furthest on the gel.
Conclusion
Mitochondria were isolated from rabbit liver sample using gradient centrifugation. The mitochondrial marker, succinate dehydrogenase, was tested in all the sample fractions by assaying for absorbance of the sample fractions using a spectrophotometer. The sample fraction proteins were analysed using SDS-PAGE. The results from the experiment proved that the most prevalent enzyme in mitochondrial fraction of a eukaryotic cell is succinate dehydrogenase.
Suggestions
For future improvement of this type of experiment caution must be taken not to introduce any contamination to the samples. This will ensure that the right results will be obtained and the right conclusions made. It is also suggested that two samples from two mammals be compared simultaneously to note and, if need be, account for any differences in their amounts of succinate dehydrogenase and other proteins.
Questions and their answers
- Are the results what you expect? If not, why?
- Yes, these are the results I expected.
- Are there any problems with the test?
- There were no problems with the test.
- What conclusions do you draw from this experiment?
- Conclusion was that the MITO fraction had the highest amount of succinate dehydrogenase because it showed the highest absorbance.
- How would you improve the strength of the test?
- The strength of the test would be improved by ensuring higher purity of the sample fractions.
- When you consider how we obtained each fraction, what subcellular components contribute to the various polypeptides you see in each lane?
- WCL lane- all cell components
- SN1 lane- microsomes
- SN2 lane- ribosomes
- MITO lane- lysosomes and peroxisomes
- Are there any bands that are unique to or especially strong in the mitochondrial fraction compared to the SN fractions? If so, what sizes are they?
- The 150 kD band was unique to the mitochondrial fraction of the sample fractions. Among all the fractions it was strongest in this fraction.
- What is a western blot? Would a western blot help you to analyse your PAGE results? What results would you expect from your experiment?
- A western blot is a technique for testing for proteins using specific antibodies. A western blot uses a nitrocellulose membrane with labelled antibodies. When antibodies bind to their corresponding proteins, a signal is given as a result of the labels fluorescing. A western blot would help analyse the PAGE results because specific antibodies would be used against the target proteins. Results from a western blot would be more specific because the technique gives signals when proteins bind their corresponding antibodies.
Reference list
Ferrer, I., 2009. Altered Mitochondria, Energy Metabolism, Voltage-Dependent Anion Channel, and Lipid Rafts Converge to Exhaust Neurons in Alzheimer’s Disease. Journal of Bioenergetics and Biomembranes, 41(5), pp.425-431.
Gray, M. W., 2012. Mitochondrial Evolution. Cold Spring Harbor Perspectives in Biology, 4(9), pp.1-17.
Murphy, M., 2009. How Mitochondria Produce Reactive Oxygen Species. Biochem. J, 417(1), pp.1-13.
Schägger, H., 1995. Native Electrophoresis for Isolation of Mitochondrial Oxidative Phosphorylation Protein Complexes. Methods in Enzymology, 260(1), p.190.
Speers, A. E., & Cravatt, B. F., 2004. Profiling Enzyme Activities in vivo Using Click Chemistry Methods. Chemistry & Biology, 11(4), pp.535-546.