Introduction
Metallothioneins (MTs)
Metallothionein (MT) proteins are characterized as low molecular weight, cysteine-rich, metal-binding proteins. The first MT was discovered more than 50 years ago as Cd2+-binding proteins present in horse kidney. Since then, the genes encoding MT proteins have been identified in the animal and plant kingdom as well as in prokaryotes like Synechococcus sp. Due to such diversity in occurrence, their function is still a debatable issue, yet it is unequivocally proven that the MTs are involved in homeostasis and transport of physiologically essential metals ions viz. Zn2+ and Cu1+ and detoxification/sequestration of toxic metal ions like Cd2+, Hg2+ etc. Besides, their role in protection against oxidative stress, regulation of cell proliferation and apoptosis, and maintenance of cellular redox balance is also well evident.
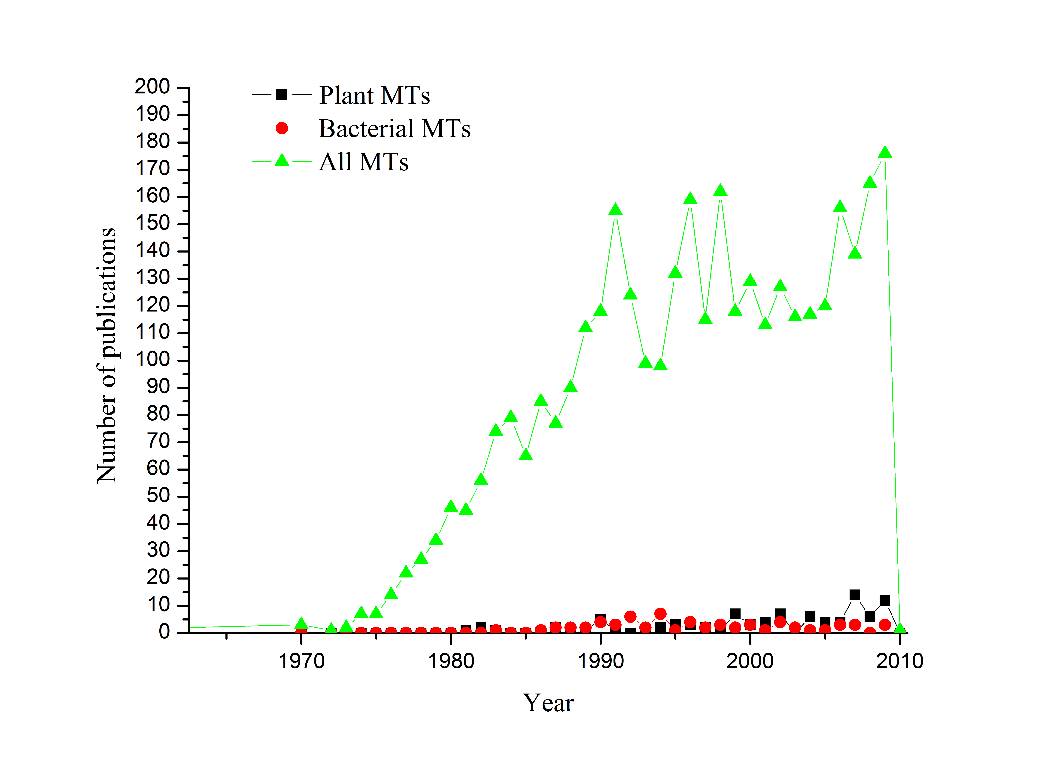
In a recent search using ScienceDirect, it was found that the number of publications under key word “Metallothionein” each year has increased from 1962 to late 2009, totaling 3494 scientific papers in this subject. Looking at the trend (Figure 1), it is clear that the interest in metallothionein has remarkably increased after eighties, and more recently between 2006 and 2009, the publication average recorded more than three papers per week. Further, it is quite clear from the figure that the interest in plant and bacterial MTs has been rather modest as compared to the overall metallothionein research field. But the trend showing an increased researched activity was clearly discernable in the figure. Understandably, the major share of the publication has come out of the work in animals and human. One of the likely reasons towards animal bias was that MTs were associated with diseases related to metal toxicity in human.
Another reason for relatively less number of publications was more to do with technical reason of the difficulties in isolation of MTs from plants and bacteria. Besides this, as reported in earlier studies the proteins were hard to be cleaved from the recombinantly expressed proteins in E. coli by proteolytic digestion. With an advent of time these problems were sorted out by way of application of fusion proteins for protein overexpression, and concomitantly more information on the basic properties of plant and bacterial MTs started to gather. The three dimensional structural data of MTs was also exclusively available from animal sources, and the recent development of structural characterization using techniques of NMR, crystallography and mass spectrometry has given adequate insight in the structure of plant and bacterial MTs.
Animal MTs have implications in inherited and occupational diseases that are linked to metabolic disorders in the handling of metal ions. Interestingly, MTs also have relevance in diagnosis of certain metal toxicosis disorders from the fact that they can serve as biomarkers, and in some cases even in cure (X-reference). The classical examples of the diseases are Menkes and Wilson’s disease (Table 1). Menkes disease is a fatal inherited X-chromosome linked recessive disorder of ATP7A gene that codes for P-type ATPase, responsible for excretion of copper and processing of certain enzymes in trans-golgi network by delivering essential prosthetic group, copper. As a result of mutation in the copper membrane protein, copper distribution to most cells in the body gets perturbed. While in most cells, systemic copper accumulation initiates neuronal degeneration and connective tissue malfunctions, in small intestine and kidneys this is manifested as deficiency in copper causing an over-expression of hepatic MT(XDaniel et al. 2004). Consequently, this copper deficiency reduces the activity of essential copper-containing enzymes, e.g. lysyl oxidase.
Another inherited disorder affecting copper homeostasis is Wilson’s disease characterized by a mutation in autosomal ATP7B gene, which codes for ATP7B that is another copper excretory membrane protein transporting copper via bile (XDaniel et al. 2004). The protein is also responsible for serum ceruloplasmin formation. The patients suffering from Wilson’s disease have complications of a failure in excretion of excess copper in bile, that lead to accumulation of copper in the liver, nervous system and other organs, and this manifest in liver failure and neurological ailments. This is despite the fact that a fraction of accumulated copper in the tissues is removed by the action of MTs.
A number of diseases caused from disturbed metal homeostasis can be occupational and can result from prolong exposure to some metals like cadmium. Currently, much attention is paid on an interesting research directed towards MTs and their potential role in neurodegenerative disorders. It has been discovered that several molecular pathways aiming to neuroprotection and regeneration are co-ordinated and often MT-mediated. For example, MTs expressed in astrocytes after CNS injury are reported to serve both neuroprotective and neuroregenerative roles critical for the outcome of recovery. Table 1 lists some common diseases associated with perturbed metal homeostasis and MTs.
Table 1. Metallothionein and disease.
a Grown inhibitory factor, MT-3, bCentral Nervous System.
General aspects of MT structures and metallation
A large number of Cys residues of MTs via mercaptide bonds enable the enzyme to bind to a variety of metal ions totaling 18 that include metals of group 11 and 12 (Zn2+, Hg2+, Ag1+, Au1+, Cd2+, Cu1+, Bi3+) and also Fe2+, Co2+, and Pb2+(X-reference). MTs exhibit preferential binding for soft metal ions specifically having the electronic configuration d10, which in case of divalent metal ions are bound in tetrahedral tetrathiolate coordination spatial geometry(X-reference). Accordingly, the UV and Circular dichroism spectra of MTs show the characteristic metal ion dependent optical features of metal–thiolate complexes as conspicuous or inconspicuous shoulders of the peptide backbone transitions towards lower energy.
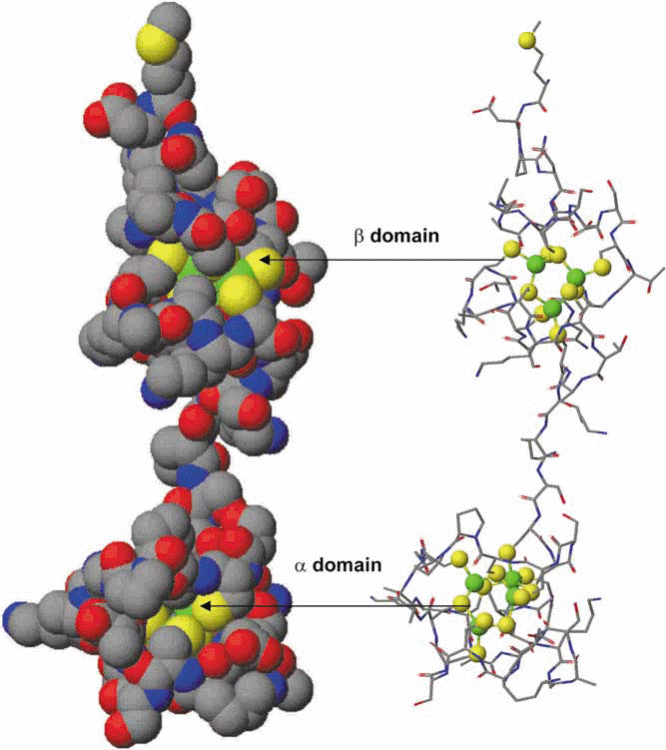
Mammalian MTs are categorized as MT-1 to MT-4 (previously MT-I to MT-IV). The typical tertiary structure of the mammalian protein consists of two domains, the N-terminal α- and C-terminal β- domains consisting of 4 and 3 metal ions, respectively, attached by a short interdomain linker sequence, resulting in a dumbbell-shaped appearance (Figure 2). The three-dimensional structures formed not only depend on the amino acid sequence, but also on the number of metal ions bound to the protein, and the coordination preference of the metal ion, i.e. tetrahedral, trigonal or digonal. Metallation reactions of the apo-MT (metal-free) and metal substitution reactions in Zn7MT can provide data about the folding mechanism of this unique protein. The mechanism by which a polypeptide chain folds to reach its native and functional structure has remained an intriguing problem of fundamental interest.
The apo-MT very quickly acquires the metals (total 7 in 4 and 3 clusters), and it is believed that metallation reaction is cooperative in nature, which indicates that binding of one metal ion accelerates affinity of the others(X-reference). As most of the tested metal ions bind to MTs in milliseconds making it difficult to work on kinetics, in a recent investigation As3+ binding kinetics was examined with apo-MT holoenzyme and its apo-α- and β- domains using ESI-MS techniques, as it takes much longer time for As3+ to bind (Ngu & Stillman, 2009). It was revealed that partially metallated intermediates exist and the binding is non-cooperative in nature. The tertiary structure of plant and bacterial MTs differ from those of mammalian due to the different Cys distribution patterns and their location within the protein sequence. However, the main driving force for folding remains the formation of the metal clusters.
Classification of MTs
MT proteins were initially classified based on the sequence and arrangement of cysteine residues. Based on this classification, Class I MTs were widely found in vertebrates including mammals, and contained 20 highly conserved cysteine residues. MTs without this strict arrangement of cysteines were referred to as Class II MTs and this included the plant MTs. In this classical MT classification system, phytochelatins (PCs) were described as Class III MTs.
Shortly after the discovery of phytochelatins as an important metal ligand required for Cd2+-tolerance in plants, a MT protein was identified in wheat, called Ec-1 protein, and subsequently a number of MT genes were identified. Further classification of Class II MT proteins in plants based on amino acid sequence could be possible. In addition to the Ec-1 protein, Robinson et al. proposed presence of two additional subclasses of “MT-like” proteins in plants. The difference was based on the location of the Cys residues, which were designated types 1 and 2. Since then, the number of characterized MT proteins from different organisms has increased dramatically, and because many of them could not be assigned to either class I or II, a more rational classifications system was felt necessary to introduce.
The presently system is based on sequence similarities of amino acids and phylogenetic relationships. The diversity of MTs was apparent as the present system has divided MTs in to 15 families (Table 2). Family 15 is comprised of the plant MTs, which have been further categorized into 4 types, namely p1, p2, p3, and pec, depending on the sequence arrangement and distribution of Cys residues (Figure 3). Family 14 encompasses the prokaryotic MTs that include as an example the MT, SmtA, from Synechococcus sp.
Table 1. Current classification of Metallothionein. In purple is the prokaryote family and in green the plant family.

Regulation of MT coding genes
One explanation as to why MTs exert a strong regulation in metabolism of diverse metal ions is that MT gene expression is generally induced by the metal ion load. MTs are coded by a family of short sequenced genes. These genes carry an upstream metal response element (MRE) that is a short DNA sequence motif of a core consensus sequence TGCRCNC (Balamurugan & Schaffner, 2009), which is a regulatory element that induces transcriptional expression of the protein in the presence of metal ions. Although metal ions are the major regulators of MT gene expression, there exist other inducers, including some glucocoticoids, cytokines, growth factors, tumor promoters etc, as well as UV radiation and physical stress.
- In mammals, metal-induced transcription of MT genes is mediated by the interaction of metal regulatory elements (MREs) with MRE-binding transcription factors, such as MTF-1, and MRE-binding factors (MRE-BFs) in the promoter region of MT genes. According to Balamurugan & Schaffner (2009), MTF-1 responds to elevated zinc levels through six highly conserved zinc-finger regions that bind to the DNA element enabling a positive regulation on MT gene expression. Apart from this, the transcriptional factor also possess cysteine region for binding with zinc. There is also another response of the heavy metals on MTF-1. Under unstressed conditions MTF-1 is confined to cytoplasm but at onset of metal load, it tends to transport inside nucleus through another conserved nuclear transport signal sequence, where it accelerates the gene expression(X-reference). In yeast, Saccharomyces cerevisiae, the interaction with copper ions activates the transcription factors Ace1 and MREs, which bind to the promoter region and in turn up-regulate the MT genes such as the CUP1 and CRS5(X-reference).
- An array of plant MT genes has been identified in angiosperms. Currently, emphasis is on plant MT gene promoters so that it may be possible to confer metal-inducibility and consequent metal tolerance by genetic engineering of such promoters to a heterologous gene. The promoter regions of some plant MT genes were found to contain both metal and hormone regulatory elements mediating the induction and efficient expression of the proteins.
A number of plants, including A. thaliana, rice, and sugarcane, contain genes encoding all the four types of MTs. This indicates an ancient nature of the four plant MT types which existed even prior to evolutionary branching of monocots and dicots. Moreover, it is likely that a majority of flowering plants also contain the four different MT types (x-reference). The presence of four types of MTs in plants with distinct arrangements of cysteines contrasts what is found in animals. For example, the four mouse MTs contain the same conserved cysteines with difference only in tissue expression. The diversity of the plant MT gene family clearly indicates not only sequence differences but also functional differences in the MT types.
Despite these assumptions, relatively much less information is available on MT genes in non-flowering plants. Notwithstanding, genes encoding p3 MTs have been cloned from several gymnosperms. Most of the algal investigations are confined to the phytochelatins in relation to heavy metal detoxification (Perales-Vela et al. 2006). A Cd2+-responsive MT-encoding gene has been isolated from Fucus vesiculocus, a brown alga, though this MT does show any sequence identity with any of the four plant type MTs described above. However, on the basis of cysteine residues, it matches with Arabidopsis MT1a and the oyster MT.
Quite like the animal counterparts, expression of plant MT genes can be induced by a range of different stimulants, which include metal ions such as Zn2+, Cu2+, and Cd2+, and phytohormones, cytokines, as well as environmental factors like low temperature, heat shock, nutrient deprivation, UV exposure etc [36-39]. In this context, MT gene expression has also been ascribed to the processes related to apoptosis, which includes leaf abscission, and the hypersensitive response towards pathogens. Occasionally, multiple forms of MTs from one subgroup can be found in one plant, displaying distinct expression patterns. For example, in A. thaliana, MT2a is highly expressed upon exposure to copper in form of CuSO4, whereas under same conditions MT2b mRNA only shows a slight increase in expression. Upon metal treatment, some plant MT genes may be regulated at the translational or even at post-translational level.
In cyanobacteria the quest of investigation was whether MTs are similar to the higher plants as they share features with eukaryotic algae, or being prokaryotic organisms are related to the bacterial type of enzymes. A line of investigation revealed that zinc-dependent expression of a well characterized metallothionein, called SmtA, in Synechococcus PCC 7942 is regulated at transcriptional leve1 by an action of the zinc sensor protein, SmtB. It was shown that SmtB, in the absence of zinc, binds to the operator-promoter region of smtA gene and represses its own transcription as well as of smtA. The operator promoter region of the smt operon (Figure 4) consists of 100 base pairs and contains two 12-2-12 imperfect inverted repeats with the core sequence 5’TGAAxx-xx-xxTTCA3’, each of which appears to be capable of binding to at least two apo-SmtB homodimers. The metal ion composition of a native MT is, to a large extent, dependents on its natural source and its previous exposure to metal ions.
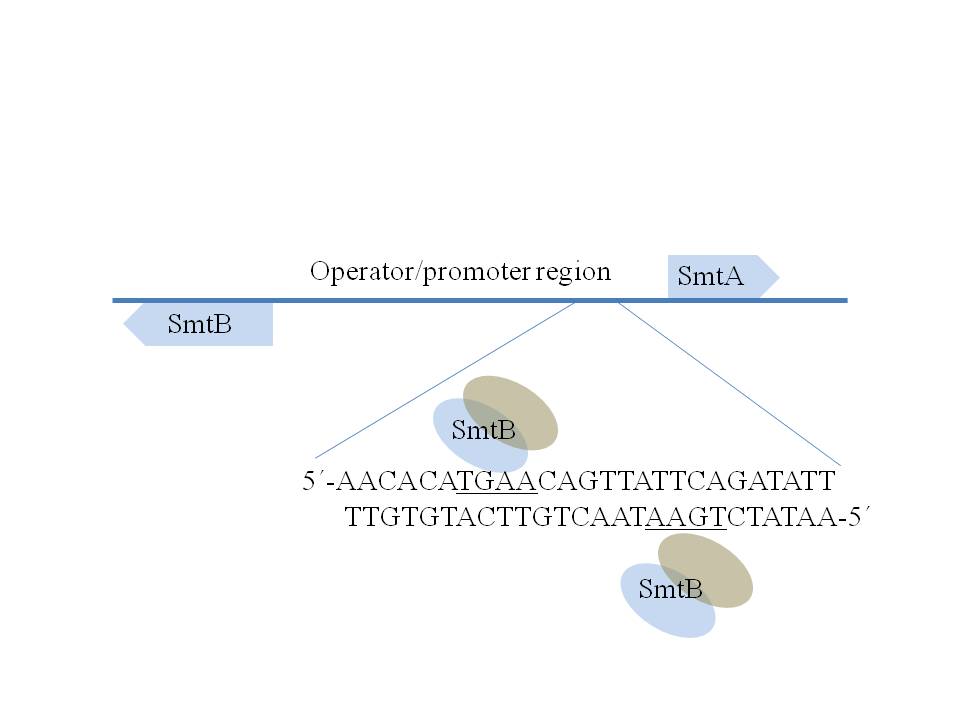
Further analysis pointed out that only one of the inverted repeats, comprising the SmtB binding sites SI and S2, appears to be sufficient for regulation of smtA expression, as depicted in Figure 4. Although SmtB homodimers were thought to have in all four zinc binding sites, only two mol equivalence of Zn2+ per dimmer seem to be sufficient to bring down its DNA affinity by negative allosteric control to a level of three order of magnitude. An analysis of a knock-out mutant of Synechococcus PCC 7942 revealed that SmtB might not be the only factor regulating smtA transcription: the cells devoid of functional smtB showed constitutive expression of SmtA even in absence of metal load, and yet an increase in smtA transcripts was observed once cells were exposed to l.4 μM of Cd2+.
The precise molecule or the mechanism behind this metalloregulation, in the absence of SmtB, remains to be investigated. It is noteworthy that a close orthologue of SmtB, termed ZiaR, regulates the expression of a zinc efflux pump (ZiaA) in another freshwater cyanobacterium, Synechocystis PCC 6803, which has been characterized as a strain devoid of metallothionein. Further, SmtB-related sensors, termed BxmR and AztR, have been characterized in Oscillaloria brevis and Anabaena PCC 7120, respectively. While BxmR is shown to regulate the expression of both efflux pump protein, Bxa1, and metallothionein, BmtA, under the control of Zn2+, Cd2+, Cu1+ and Ag1+, the AztR protein was shown to exclusively regulate the expression of a highly zinc- specific efflux pump mediated through AztA protein. A rice field cyanobacterium of genus Anabaena also contains a metallothionein, but not much information is available on its regulation.
More recently, structures, mechanisms, and phylogeny of SmtB/ArsR-like sensors have been discussed in greater detail. The ars operon comprising arsRDABC cistrons in gram positive and negative bacteria brings about arsenic resistance. In E. coli this is a plasmid R773-borne operon with arsR being the arsenic inducible repressor (Diorio et al. 1995). The metalloregulatory proteins belonging to the ArsR family respond to not only arsenic but also to antimony, cadmium and zinc. Leaving these few examples, it should be emphasized that a majority of cyanobacterial genome sequences do not contain putative SmtB orthologues, even though the sequences representing the genes for bacterial MTs may be present. This suggests that there exist a large number of metal ion homeostasis mechanisms in cyanobacteria, which include different degrees of specificity of metal-mobilizing genes. Clearly, the complete understanding of MT gene regulation in bacteria deserves further attention.
Metallothioneins in plants
While the discovery of the first mammalian metallothioneins dates back more than 50 years, the first cysteine-rich small protein from plant material was isolated only 25 years ago from wheat (Triticum aestivum) germs. This protein was shown to be solely encoded by the mRNA, preserved in the dry mature of wheat embryos, already started to decline around 5 hours past seed imbibition. Recognizing a very high content (20-25%) of total cysteine present in the nascent proteins as compared to the average cellular proteins, it was denoted as the “early cysteine-labeled protein” or Ec. Originally Ec was isolated as S-alkylated protein following iodoacetamide treatment of wheat germ soluble protein fraction to a yield of 5-10 mg/ 100 g of plant material. The unique amino acid composition, without having any prior information on the sequence or on its ability to bind to certain metal ions, has given enough impetus to establish a relationship of the Ec with the similar Cys-rich mammalian MTs.
However, it was the information on partial amino acid sequence, the metal ion content, and the spectrophotometric data of the Cd2+-substituted T. aestivum Ec as the first metallothionein that has triggered a new line of investigation in higher plants that was substantiated later by sufficient evidences. Apart from the presence of His residues, Ec actually met the other criteria justifying the designation of Cys- and metal ion-rich proteins as metallothioneins, such as: low molecular weight (Mr = 5-10 kDa), the absence of aromatic amino acids (other than His), a characteristic sequence pattern of Cys residues like “CysXaaCys” motifs with Xaa representing an amino acid other than Cys, and the characteristic optical spectra of metal-thiolate complexes with discernable shoulder of peptide backbone transitions around 250nm for the Cd2+-substituted forms.
Since this early finding, the number of newly discovered plant MT sequences, determined mostly from their mRNA level, has substantially increased. For example, a search in the Swiss-Prot protein sequence database using the keyword “metallothionein” currently lists 259 entries, and 64 of these sequences originate from the plant kingdom. In addition, recently the first tertiary structure of a plant metallothionein was determined (Figure 5). For this, the C-terminal βE-Ec-1 domain was derived from proteolytic digestion of Triticum aestivum Ec-1 MT protein with the help of bacterial proteinase K.
When comparing the plant MTs to the better characterized vertebrate isoforms, two characteristic distinguishing features became obvious. First, plant MTs may contain the aromatic amino acids, phenylalanine, tyrosine and histidine, which are strictly absent in the vertebrate counterparts. Secondly, in the amino acid sequences of plant MTs, 10 to 45 non-cysteine residues separate the two regions where the cysteines are concentrated (Figure 3). In contrast, the cysteine-rich regions of the vertebrate MTs are separated by only two to four residues, predominantly lysines. Even within the plant MTs, a vast diversity of amino acid sequences as well as differences in the number and arrangement of cysteines residues was found, suggesting that most likely different biological function(s) are assigned to different MT subfamilies. The variation between the plant and the vertebrate forms and within the plant sub-forms gave sufficient indication towards differences in physiological properties, recognizing the importance of plant MTs for future investigations.
In MTs belonging to subfamily p1, 12 cysteine residues are arranged exclusively in Cys-Xxx-Cys motifs equally divided between two cysteine-rich regions (Figure 3) and separated from each other by about 40 amino acids. A variant, also belonging to subfamily p1, is called p21 that includes proteins containing one additional cysteine residue in the C-terminal as well as a short linker composed of seven amino acids. Typically, p2 MTs also have two cysteine-rich domains separated by about 40 residues. A majority of their 14 cysteines are arranged in Cys-Xxx-Cys motifs, however, in the N-terminal domain, four of them are arranged in Cys-Cys and Cys-Xxx-Xxx-Cys motifs. Subfamily p3 MTs feature total ten cysteines that are grouped in two regions separated by about 35 amino acids. At the N-terminal there are four cysteines, out of which only two forms a Cys-Xxx-Cys motif.
The pec subfamily has the highest cysteine content of all the plant MTs. 17 cysteines are located in three distinct regions with six (N-terminal and central cysteine-rich regions) or five (C-terminal cysteine-rich region) conserved cysteines separated by 10 and 15 residues. The central cysteine-rich domain shows an identical conservation pattern to that of the C-terminal regions of the members of p1, p2 and p3 subfamilies. Besides, two histidine residues in the central region are also highly conserved within the pec subfamily. Despite the present knowledge of amino acid sequences, only a single three-dimensional structure, the one of βE-Ec-1, is known until now, yet it provides enough details on the possible structures of the subfamilies of plant MTs. It shows a cluster arrangement that was not reported for any metallothionein so far analysed, reiterating to the fact that plant MTs might have different hitherto unknown functions compared to the mammalian forms.
Musa acuminata MT3
M. acuminata MT3 belongs to the plant MT fruit-specific p3 subfamily. Having a total of 65 amino acids (Figure 6) including 10 cysteine and 1 histidine residues, it features a cysteine content and percentage that is more comparable to fungal and prokaryotic MTs rather than the mammalian forms. Additionally, as per the members of the p3 subfamily, it shows characteristic long Cys-free linker region between the Cys-rich regions.

The mRNA transcripts of MT3 genes are most abundant in ripening fleshy fruits, but are barely detectable in the earlier stages of fruit development or in flowers. MT3 is most likely not directly involved in ripening processes which involve cell wall softening, and sugar and pigment accumulation. Here, MT3 can be assumed to have a role in combating the stress conditions, because like the other oxidative stress related genes found in ripening fruits, MT3 might also exhibit a similar function. A role in scavenging excess of Zn2+ released upon protein metabolism is also conceivable, and in this way MT3 could be employed as a reservoir of this essential element for future use for fruit ripening. This is in analogy to the proposed function of MTs in the senescencing tissues. High MT3 mRNA levels were also detected both in green and senescent leaves of buckwheat, and can be up-regulated by addition of Zn2+ or Cu2+ ions.
The metal content of plant MTs is believed to be lower than in mammalian MTs, but to make a statement sufficient information is lacking. For the p3 subfamily, only two protein species have been investigated in detail. A modeling study was performed with Actinidia chinensis (Kiwi) MT3, a member of the p3 subfamily devoid of the eleventh coordinating residue that is a Cys or His, and this has resulted in a theoretical value of 3 Cd2+ ions per protein on the basis of two domain structure. The purified GST-MT3 fusion protein of Elaeis guineensis MT3, which is closely related to M. acuminata MT3, also features an additional His residue. This protein was found to bind an additional amount of 1.7 Zn2+ ions when compared to the GST-fusion protein alone. Thus, from these differences it is obvious that more precise information on MTs belonging to this subfamily is desirous. Therefore, this thesis aims to provide data to allow a better understanding of the properties of these proteins.
Metallothioneins in bacteria
The occurrence of MTs was initially thought to be restricted to only the eukaryotic phyla. Later studies revealed the presence of MT gene homologues and orthologues in the genome sequences of cyanobacteria and proteobacteria such as the pseudomonads and Staphylococcus epidermidis (Figure 7). Even functionally, the existence of MTs was quite evident. Phenotyping for the SmtA gene of Synechococcus PCC7942 has clearly shown that transcription is induced by zinc load, and that deletion of the SmtA gene results in a substantial alleviation of a capability to tolerate high concentrations of zinc. Apart from the gram negative bacteria, recently copper regulated MT genes, termed mymT, have been sequenced from pathogenic gram positive species of Mycobacteria (Gold et al. 2008). Interestingly, MymT displays little homology with the existing prokaryotic and eukaryotic MTs. The first Cys rich motif of MymT (Cys-X-His-X-X-Cys-X-Cys-) presents similarity to SmtA but the second Cys rich motif (Cys-His-Cys-X(2)-Cys-X(2)-Tyr-Arg-Cys-Thr-Cys-) has practically no analogy to any corresponding regions from any organism.
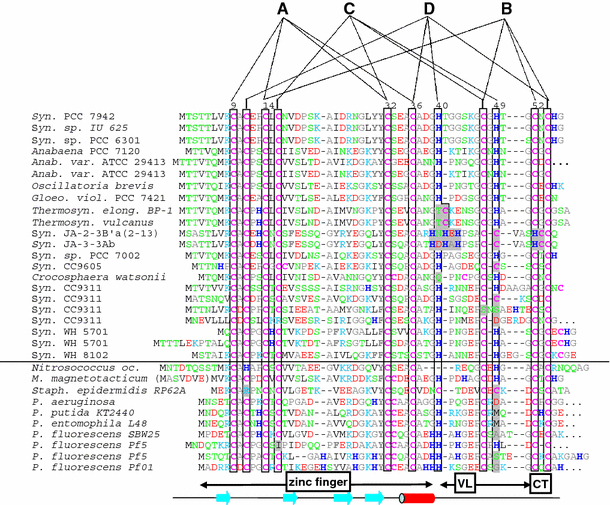
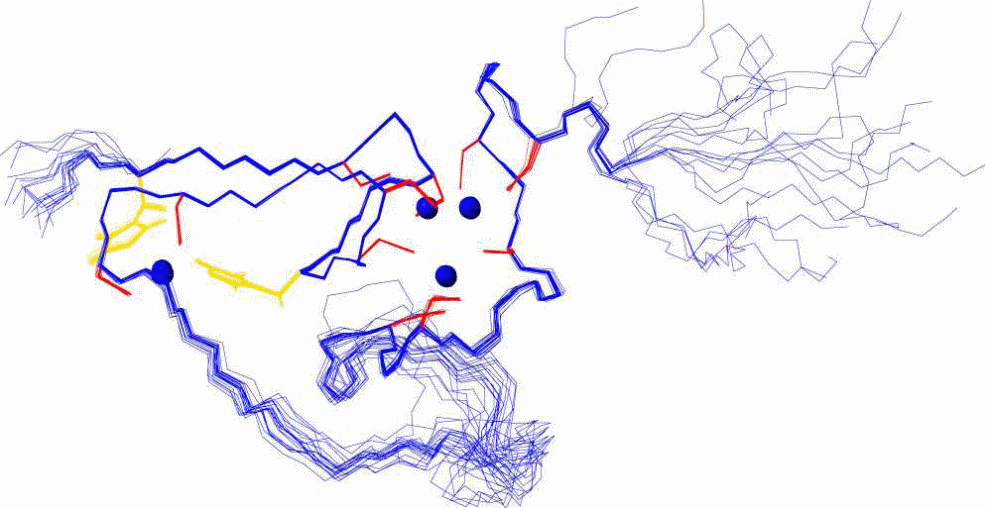
Since SmtA was the only prokaryotic protein available, the structure, solution properties, and metal-binding dynamics of this protein has been elaborately characterized, and based on several properties it was recognized as a different protein from the other zinc-binding eukaryotic MTs. For example, mammalian MT-1 and MT-2 are two-domain proteins and bind zinc (or cadmium) ions in two metal–sulfur clusters with compositions M3Cys9 (β domain) and M4Cys11 (β domain). Apart from β-turns, the folded proteins contain virtually no regular secondary structural elements, and therefore the structure is considered to be dominated by the metal clusters. In contrast, the zinc-binding SmtA from Synechococcus PCC 7942 consists of only one domain containing three- or four metal clusters, and one of the zinc ions (labeled A in Figure 8) forms part of a treble-clef zinc finger-like fold.
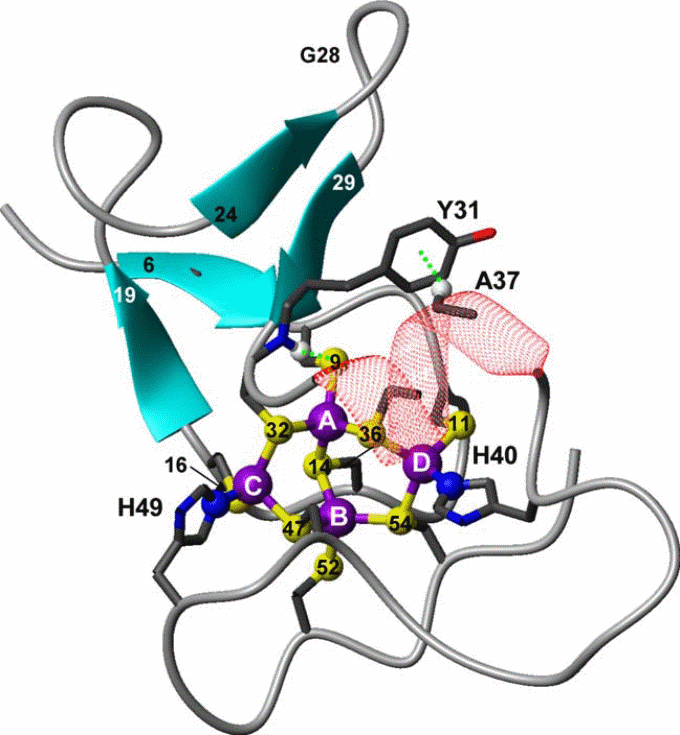
In the cyanobacterial MT SmtA, the protein fold is further stabilized by a CH–π interaction between the aromatic side chain of a tyrosine residue from the hairpin and an alanine CH(α) from the helix (Figure 8). However, the most distinctive bioinorganic feature is the participation of histidine residues in zinc coordination. Taking an example of SmtA and MymT it is apparent that histidine, aspartate and methionine can coordinate in metal binding, substituting cysteine in several bacteria (x-reference). This is also another distinguishing feature from the mammalian MTs. The MT-1, MT-2, and MT-3, all 20 sulfur ligands in the two clusters, are provided by Cys residues from the protein chain.
In comparison, the metal cluster in Synechococcus SmtA consists of four Zn2+ ions, coordinating with nine Cys thiolate sulfurs, and two His imidazole nitrogen. Despite the substitution of two cysteines by histidines, the structure of the cluster in SmtA is remarkably similar to that of the native M4Cys11 cluster in the mammalian MT α-domains. Although mixed cysteine–histidine coordination of zinc is a novel feature for MTs, especially in bacteria, it is common among zinc fingers, including the classical Cys2His2 finger . In SmtA, the histidine residues are not ligands for the ‘‘finger’’ zinc, but for two adjacent zinc ions in the cluster (Figure 8). In zinc-finger proteins, mutation to cysteine/histidine residues can have drastic change in protein folding; for example, in retroviral nucleocapsid proteins, in which the Cys3His motif is replaced by either a Cys4 or a Cys2His2 motif, the mutants displayed completely different protein conformations and the corresponding viruses turned avirulent, despite the fact that zinc was still bound.
Although Cys and His are the two prevalent zinc-binding residues in “Zn finger” proteins, there is surprisingly little information on the thermodynamics and kinetics of zinc binding to the corresponding sites with prevalent residues. Furthermore, our understanding as to why certain coordination environment for a metal is favored over others is still redundant.
Synechococcus vulcanus MT (SvulcMT)
The metallothionein SvulcMT is reported from a thermophilic cyanobacterium, Synechococcus vulcanus, and it contains net 55 amino acids (Figure 9), of which 10 are Cys and 2 His residues. So far, there is no information available with respect to its metal ion binding capacity and stability, and its participation in metal homeostasis.

Thermophiles inhabit some of the planet’s harshest environments such as boiling deep-sea rift vents, volcanic craters and polar glaciers. Cyanobacteria are ubiquitous for their presence in such environments, owing to some unique genetic adaptations that protect them. For example, the heat tolerance among thermophilic microorganisms is conferred by variation in protein and nucleic acid configurations that are less reliant on hydrogen bonding to preserve their spatial structure.
The hot spring mat communities in terrestrial habitats have also attracted attention of scientists because of unique adaptations of the microbial flora to the harsh environments. Since these harsh conditions hardly allow animal growth, an absence of predators considerably change the food web in these ecosystems, and microbial diversity and community structure is principally determined by temperature and salinity gradients prevailing in the spring systems, or by changing water chemistry between the spring systems. An example of species diversity in cyanobacterial community is on account of invasion of members of Synechococcus in alkaline hot spring habitats across temperate waters (Western North America, Asia, Africa, and Europe). Hot spring outflows bring remarkable temperature gradients, and microbial communities with dominant Synechococcus generally thrive in these ecosystems, in which temperatures vary between ~45°C and 73°C. Incidentally, this is the thermal maxima for photosynthetic life. The thermophilic cyanobacterium, S. vulcanus, an inhabitant of such hot springs, is characterized by an optimum growth temperature of 55 ºC.
Although metallothionein (MT)-‘like’ metal-ligands have been reported in prokaryotes, MTs from only a few bacteria has been isolated and sequenced. One can assume profound difference in MTs of thermophilic micro-organisms from other species. This could be one of the explanations for no apparent similarity between the amino acid sequence of SvulcMT protein and mammalian MTs. The amount of SvulcMT in S. vulcanus cells was found to increase after supplementing Zn2+ or Cd2+, but not Cu2+ in the growth medium. The isolated native protein was found to anchor predominantly Zn2+ or Cd2+ depending on the metal supplemented in the medium. In addition, small amount of Cu2+ was also detected upon induction with Cd2+ or Zn2+. Analysis of the native Zn2+-protein partially resembled to the mammalian Zn-MT. However, the metal affinities and conformation of the metal-binding site(s) is still elusive. It remains to be seen that whether analogous to the eukaryotic MTs this prokaryotic protein has equivalent high affinity for metals.
Possible application of metallothioneins
Metallothionein is a protein that demands further research, and this has attracted scientists of different disciplines. The state of present knowledge projects that future research on metallothionein is expected to contribute towards understanding metal associated biological effects with some application potential. A possible application that can be thought is incorporation of MTs as biosensors. A biosensor encompasses coupling of a biological molecule like an enzyme with a microelectronic device to help detection of an analyte in a given sample. Several bacterial proteins including metallothioneins conferring metal resistance has implications in developing sensory devices for environmental monitoring.
This strategy can be extended to even some other organisms. Fundamental requirements for the applicability of certain protein as biosensor are: high stability, a capability to reversibly bind the metal ions, and specificity. Theoretically, metallothioneins can fulfil at least the first two criteria, but with present understanding they are not expected to discriminate between different soft metal ions. The interaction between heavy metals and SmtA, immobilized on gold as a GST-fusion protein, elicited capacitance changes that allows detection of fentomolar to millimolar concentrations of Hg2+, Cu2+, Zn2+, and Cd2+ in pure solutions. Although the responses to different metals are not identical, specific metal detection was not an easy preposition with SmtA. It is an intriguing perspective to make use of the zinc finger scaffold and employ site-directed mutagenesis to improvise specificity towards a given metal.
Another far reaching though interesting application is the use of MT in bioremediation of heavy metals. The remediation of anthropogenic heavy metals is a serious concern and differs from degradation of the organic pollutants to harmless chemical substances taking help of bacteria. In most instances heavy metal remediation is only possible by immobilization and isolation of the metals in non-bioavailable form, sometimes enabling biomining. The observation that bacteria can synthesize metallothioneins with capability of intracellular metal accumulation has opened a sector for application of bacteria as metal bioaccumulators to remediate the heavy-metal polluted soil, sediments, and wastewater.
In a real sense, cadmium-resistant pseudomonads (P. putida and P. fluorescens) can be isolated directly from sewage sludge, even though state of-the-art of genetic engineering promises an easier preposition for the construction of strains with a tailored repertoire. Most studies aiming at the use of recombinant metallothioneins expressed in bacteria for bioremediation involve the heterologous expression of eukaryotic MTs, including the fusion of MTs to outer membrane proteins to enhance bio-adsorption. Sode et al. demonstrated that insertion of the SmtA gene from freshwater Synechococcus PCC 7942 into a zinc-sensitive, SmtA-inactivated marine Synechococcus strain NKBG IS04Ic has led to an elevated tolerance towards zinc, copper, and cadmium, and suggested that such strains could be of use for bioremediation in the marine environments.
References
P.-A. Binz, J.H.R. Kägi, Metallothionein: Molecular evolution and classification, in: C. Klaassen (Ed.), Metallothionein IV, Birkhäuser Verlag, Basel, 1999, pp. 7-13.
W. Maret, The Function of Zinc Metallothionein: A Link between Cellular Zinc and Redox State, J. Nutr. 130 (2000) 1455S-1458S.
R.D. Palmiter, Proc. Natl. Acad. Sci. USA 95 (1998) 8428.
W. Braun, M. Vašák, A.H. Robbins, C.D. Stout, G. Wagner, J.H.R. Kägi, K. Wüthrich, Comparison of the NMR solution structure and the X-ray crystal-structure of rat metallothionein-2, Proc. Natl. Acad. Sci. U S A 89 (1992) 10124-10128.
F. Reinecke, O. Levanets, Y. Olivier, R. Louw, B. Semete, A. Grobler, J. Hidalgo, J. Smeitink, A. Olckers, F.H. Van der Westhuizen, Metallothionein isoform 2A expression is inducible and protects against ROS-mediated cell death in rotenone-treated HeLa cells, Biochem. J. 395 (2006) 405-415.
G. Nordberg, T. Jin, P. Leffler, M. Svensson, T. Zhou, M. Nordberg, Metallothioneins and diseases with special reference to cadmium poisoning, Analusis 28 (2000) 396-400.
J.G. Hoey, S.H. Garrett, M.A. Sens, J.H. Todd, D.A. Sens, Expression of MT-3 mRNA in human kidney, proximal tubule cell cultures, and renal cell carcinoma, Toxicology Letters 92 (1997) 149-160.
R.S. Chung, J. Hidalgo, A.K. West, New insight into the molecular pathways of metallothionein-mediated neuroprotection and regeneration, Journal of Neurochemistry 104 (2008) 14-20.
E. Freisinger, Metallothioneins in plants, Met. Ions Life Sci. 5 (2009) 107-153.
J.H.R. Kägi, S.R. Himmelhoch, P.D. Whanger, J.L. Bethune, B.L. Vallee, Equine hepatic and renal metallothioneins – Purification, molecular-weight, amino-acid composition, and metal content, J. Biol. Chem. 249 (1974) 3537-3542.
Y. Kojima, C. Berger, B.L. Vallee, J.H.R. Kägi, Amino-acid sequence of equine renal metallothionein-1B, Proc. Natl. Acad. Sci. U S A 73 (1976) 3413-3417.
J. Chan, Z. Huang, M.E. Merrifield, M.T. Salgado, M.J. Stillman, Studies of metal binding reactions in metallothioneins by spectroscopic, molecular biology, and molecular modeling techniques, Coordination Chemistry Reviews 233-234 (2002) 319-339.
W. Lu, M. Kasrai, G.M. Bancroft, M.J. Stillman, K.H. Tan, Sulfur L-edge XANES study of zinc-, cadmium-, and mercury-containing metallothionein and model compounds, Inorganic Chemistry 29 (2002) 2561-2563.
H. Berg, Metallothionein (IV) C. Klaassen (Ed.); Birkhäuser, Basel, 1999, XLV+645 pages, ISBN 3-7643-5830-0, DM 298.00, Bioelectrochemistry 51 (2000) 83-83.
G.W. Stuart, P.F. Searle, R.D. Palmiter, Identification of multiple metal regulatory elements in mouse metallothionein-I promoter by assaying synthetic sequences, Nature 317 (1985) 828-831.
W.R. Bidlack, Metal Ions in Biological System, Volume 36: Interrelations Between Free Radicals and Metal Ions in Life Processes, J Am Coll Nutr 18 (1999) 368-369.
K.T. Suzuki, N. Imura, M. Kimura, Metallothionein III 1993.
M.G. Cherian, Y.J. Kang, Metallothionein and Liver Cell Regeneration, Exp. Biol. Med. 231 (2006) 138-144.
B.G. Lane, R. Kajioka, T.D. Kennedy, The wheat-germ Ec protein is a zinc-containing metallothionein, Biochem. Cell Biol. 65 (1987) 1001-1005.
N.J. Robinson, A.M. Tommey, C. Kuske, P.J. Jackson, Plant metallothioneins, Biochem. J. 295 (1993) 1-10.
A. Murphy, L. Taiz, Comparison of metallothionein gene expression and nonprotein thiols in ten Arabidopsis ecotypes. Correlation with copper tolerance, Plant Physiol. 109 (1995) 945-954.
W.E. Rauser, Phytochelatins, Annu. Rev. Biochem. 59 (1990) 61-86.
Y. Kojima, P.-A. Binz, J.H.R. Kägi, Nomenclature of metallothionein: Proposal for a revision, in: C. Klaassen (Ed.), Metallothionein IV, Birkhäuser Verlag, Basel, 1999, pp. 3-6.
C. Cobbett, P. Goldsbrough, Phytochelatins and metallothioneins: roles in heavy metal detoxification and homeostasis, Annu. Rev. Plant Biol. 53 (2002) 159-182.
E. Freisinger, Plant MTs – long neglected members of the metallothionein superfamily, Dalton Trans. (2008) 6663-6675.
G.K. Andrews, Regulation of Metallothionein Gene Expression by Oxidative Stress and Metal Ions, Biochem. Pharmacol. 59 (2000) 95-104.
A.T. Miles, G.M. Hawksworth, J.H. Beattie, V. Rodilla, Induction, regulation, degradation, and biological significance of mammalian metallothioneins, Crit. Rev. Biochem. Mol. 35 (2000) 35-70.
S. Labbe, D.J. Thiele, Pipes adn wiring: the regulation of copper uptake and distribution in yeast, Trends Miocrobiol. 7 (1999) 500-505.
I.M. Evans, L.N. Gatehouse, J.A. Gatehouse, N.J. Robinson, R.R.D. Croy, MT in pea, FEBS Lett. 262 (1990) 29-32.
C.A. Whitelaw, J.A. Le Huquet, D.A. Thurman, A.B. Tomett, The isolation and characterization of type II metallothionein-like genes from tomato (Lycopersicon esculentum L.), Plant Mol. Biol. 33 (1997) 503-511.
J. Zhou, P.B. Goldsbrough, Functional homologs of animal and fungal metallothionein genes from Arabidopsis, Plant Cell 6 (1994) 875-884.
M. Charrhai, M. Osusky, L. Osusky, D. Yevtushenko, S. Misra, Functional analysis of a Douglas-fir metallothionein-like gene promoter: transient assays in zygotic and somatic embryos and stable transformation in transgenic tobacco, Planta 220 (2004) 118-128.
C.D. Klaassen, J. Liu, S. Choudhuri, Metallothionein: An intracellular protein to protect against cadmium toxicity, Annu. Rev. Pharmacol. 39 (1999) 267-294.
M. Chatthai, K.H. Kaukinen, T.J. Tranbarger, P.K. Gupta, S. Misra, The isolation of a novel metallothionein-related cDNA expressed in somatic and zygotic embryos of Douglas-fir: regulation by ABA, osmoticum, and metal ions, Plant Molecular Biology 34 (1997) 243-254.
C.A. Morris, B. Nicolaus, V. Sampson, J.L. Harwood, P. Kille, Identification and characterization of a recombinant metallothionein protein from a marine alga, Fucus vesiculosus, Biochem. J. 338 (1999) 553-560.
H.M. Hsieh, W.K. Liu, A. chang, P.C. Huang, RNA expression patterns of a type 2 metallothionein-like gene from rice, Plant Mol. Biol. 32 (1996) 525-529.
X. Quan, H. Zhang, L. Shan, Y. Bi, Advances in plant metallothionein and its heavy metal detoxification mechanisms, Hereditas (Beijing) 28 (2006) 375-382.
P. Liu, C.J. Goh, C.S. Loh, E.C. Pua, Differential expression and characterization of three metallothionein-like genes in Cavendish banana (Musa acuminata), Physiol. Plant 114 (2002) 241-250.
S.A. Coupe, J.E. Taylor, J.A. Roberts, Characterisation of an mRNA encoding a metallothionein-like protein that accumulates during ethylene-promoted abscission of Sambucus nigra L. leaflets, Planta 197 (1995) 442-447.
A. Butt, C. Mousley, K. Morris, J. Beynon, C. Can, E. Holub, J.T. Greenberg, V. Buchanan-Wollaston, Differential expression of a senescence-enhanced metallothionein gene in Arabidopsis in response to isolates of Peronospora parasitica and Pseudomonas syringae, Plant J. 16 (1998) 209-221.
J. Zhou, P.B. Goldsbrough, Structure, organization and expression of the metallothionein gene family in Arabidopsis, Mol. Gen. Genet. 248 (1995) 318-328.
A.P. Morby, J.S. Turner, J.W. Huckle, N.J. Robinson, SmtB is a metal-dependent repressor of the cyanobacterial metallothionein gene smtA: identification of a Zn inhibited DNA-protein complex, Nucl. Acids Res. 21 (1993) 921-925.
N. Romero-Isart, M. Vašák, advances in the structure and chemistry of MTs, J. Inorg. Biochem. 88 (2002) 388-396.
C. Blindauer, Bacterial Metallothioneins, in: H.S.a.R.K.O.S. Astrid Sigel (Ed.), Metal ions in life science, vol. 5, RSC, 2009.
M.L. VanZile, X. Chen, D.P. Giedroc, Allosteric Negative Regulation of smt O/P Binding of the Zinc Sensor, SmtB, by Metal Ions: A Coupled Equilibrium Analysis†Biochemistry 41 (2002) 9776-9786.
J.S. Turner, N.J. Robinson, Cyanobacterial metallothioneins: Biochemistry and molecular genetics, Journal of Industrial Microbiology and Biotechnology 14 (1995) 119-125.
C. Thelwell, N.J. Robinson, J.S. Turner-Cavet, An SmtB-like repressor from Synechocystis PCC 6803 regulates a zinc exporter, Proceedings of the National Academy of Sciences of the United States of America 95 (1998) 10728-10733.
L. Tong, N. Susumu, H. Kazunobu, U. Yoshiko, S. Mineo, K. Maki, K. Kunihiro, A metallothionein and CPx-ATPase handle heavy-metal tolerance in the filamentous cyanobacterium Oscillatoria brevis1 1 The nucleotide sequence data reported in this paper will appear in the DDBJ, EMBL, and GenBank databases under accession numbers AB085749 (for bmtA) and AB073990 (for bxa1), FEBS Letters 542 (2003) 159-163.
T. Liu, S. Nakashima, K. Hirose, M. Shibasaka, M. Katsuhara, B. Ezaki, D.P. Giedroc, K. Kasamo, A Novel Cyanobacterial SmtB/ArsR Family Repressor Regulates the Expression of a CPx-ATPase and a Metallothionein in Response to Both Cu(I)/Ag(I) and Zn(II)/Cd(II), Journal of Biological Chemistry 279 (2004) 17810-17818.
T. Liu, J.W. Golden, D.P. Giedroc, A Zinc(II)/Lead(II)/Cadmium(II)-Inducible Operon from the Cyanobacterium Anabaena Is Regulated by AztR, an α3N ArsR/SmtB Metalloregulator†Biochemistry 44 (2005) 8673-8683.
A.B. Claudia, D.H. Mark, K.R. Andrea, A.P. John, W.B. Peter, J.S. Peter, J.R. Nigel, Multiple bacteria encode metallothioneins and SmtA-like zinc fingers, Molecular Microbiology 45 (2002) 1421-1432.
S. Tottey, D.R. Harvie, N.J. Robinson, Understanding How Cells Allocate Metals Using Metal Sensors and Metallochaperones, Accounts of Chemical Research 38 (2005) 775-783.
M.A. Pennella, D.P. Giedroc, Structural Determinants of Metal Selectivity in Prokaryotic Metal-responsive Transcriptional Regulators, BioMetals 18 (2005) 413-428.
A.B. Claudia Zinc-Handling in Cyanobacteria: An Update, Chemistry & Biodiversity 5 (2008) 1990-2013.
M. Margoshes, B.L. Vallee, A Cadmium Protein from Equine Kidney Cortex, J. Am. Chem. Soc. 79 (1957) 4813-4814.
L. Hanley-Bowdoin, B.G. Lane, A novel protein programmed by the mRNA conserved in dry wheat embryos. The principal site of cysteine incorporation during early germination, Eur. J. Biochem. 135 (1983) 9-15.
E.A. Peroza, E. Freisinger, Metal ion binding properties of Triticum aestivum Ec-1 metallothionein: Evidence supporting two separate metal-thiolate clusters, J. Biol. Inorg. Chem. 12 (2007) 377-391.
J. Domènech, G. Mir, G. Huguet, M. Capdevila, M. Molinas, S. Atrian, Plant metallothionein domains: functional insight into physiological metal binding and protein folding, Biochimie 88 (2006) 583-593.
S.E. Ledger, R.C. Gardner, Cloning and characterization of five cDNAs for genes differentially expressed during fruit development of kiwifruit (Actinidia deliciosa var.deliciosa), Plant Mol. Biol. 25 (1994) 877-886.
T. Moriguchi, M. Kita, S. Hisada, T. Endo-Inagaki, M. Omura, Characterization of gene repertoires at mature stage of citrus fruits through random sequencing and analysis of redundant metallothionein-like genes expressed during fruit development, Gene 211 (1998) 221-227.
S.J. Reid, G.S. Ross, Up-regulation of two cDNA clones encoding metallothionein-like proteins in apple fruit during cool storage, Physiol. Plant. 100 (1997) 183-189.
Y.W. Nam, L. Tichit, M. Leperlier, B. Cuerq, I. Marty, J.M. Lelievre, Isolation and characterization of mRNAs differentially expressed during ripening of wild strawberry (Fragaria vesca L.) fruits, Plant Mol. Biol. 39 (1999) 629-636.
R. Moyle, D.J. Fairbairn, J. Ripi, M. Crowe, J.R. Botella, Developing pineapple fruit has a small transcriptome dominated by metallothionein, J. Exp. Bot. 56 (2005) 101-112.
J.M. Brkljacic, J.T. Samardzic, G.S. Timotijevic, V.R. Maksimovic, Expression analysis of buckwheat (Fagopyrum esculentum Moench) metallothionein-like gene (MT3) under different stress and physiological conditions, J. Plant Physiol. 161 (2004) 741-746.
Z. Chunming, L. Tun, Z. Riqing, Z. Nanming, L. Jinyuan, Modeling of kiwifruit metallothionein kiwi503, Chinese Sci. Bull. 45 (2000) 1413-1417.
S.N.A. Abdullah, S.C. Cheah, D.J. Murphy, Isolation and characterisation of two divergent type 3 metallothioneins from oil palm, Elaeis guineensis, Plant Physiol. Bioch. 40 (2002) 255-263.
R.W. Olafson, W.D. McCubbin, C.M. Kay, Primary- and secondary-structural analysis of a unique prokaryotic metallothionein from a Synechococcus sp. cyanobacterium, Biochem. J. 251 (1988) 691-699.
J.S. Turner, A.P. Morby, B.A. Whitton, A. Gupta, N.J. Robinson, Construction of Zn2+/Cd2+ hypersensitive cyanobacterial mutants lacking a functional metallothionein locus, Journal of Biological Chemistry 268 (1993) 4494-4498.
C. Blindauer, M. Razi, D. Campopiano, P. Sadler, Histidine ligands in bacterial metallothionein enhance cluster stability, Journal of Biological Inorganic Chemistry 12 (2007) 393-405.
C.A. Blindauer, M.D. Harrison, J.A. Parkinson, A.K. Robinson, J.S. Cavet, N.J. Robinson, P.J. Sadler, A metallothionein containing a zinc finger within a four-metal cluster protects a bacterium from zinc toxicity, Proc. Natl. Acad. Sci. U S A 98 (2001) 9593-9598.
O.I. Leszczyszyn, R. Schmid, C.A. Blindauer, Toward a property/function relationship for metallothioneins: Histidine coordination and unusual cluster composition in a zinc-metallothionein from plants, Proteins 68 (2007) 922-935.
A. Arseniev, P. Schultze, E. Worgotter, W. Braun, G. Wagner, M. Vašák, J.H.R. Kägi, K. Wüthrich, Three-dimensional structure of rabbit liver [Cd7]metallothionein-2a in aqueous solution determined by nuclear magnetic resonance, J. Mol. Biol. 201 (1988) 637-657.
N.V. Grishin, Treble clef finger–a functionally diverse zinc-binding structural motif, Nucl. Acids Res. 29 (2001) 1703-1714.
C.A. Blindauer, P.J. Sadler, How to Hide Zinc in a Small Protein, Accounts of Chemical Research 38 (2004) 62-69.
C.A. Blindauer, M.D. Harrison, J.A. Parkinson, A.K. Robinson, J.S. Cavet, N.J. Robinson, P.J. Sadler, A metallothionein containing a zinc finger within a four-metal cluster protects a bacterium from zinc toxicity, Proceedings of the National Academy of Sciences of the United States of America 98 (2001) 9593-9598.
M.J. Daniels, J.S. Turner-Cavet, R. Selkirk, H. Sun, J.A. Parkinson, P.J. Sadler, N.J. Robinson, Coordination of Zn2+ (and Cd2+) by Prokaryotic Metallothionein, Journal of Biological Chemistry 273 (1998) 22957-22961.
A. Klug, D. Rhodes, Zinc fingers : a novel protein motif for nucleic acid recognition, 12 (1987) 464-469.
R.H. Stote, E. Kellenberger, H. Muller, E. Bombarda, B.P. Roques, B. Kieffer, Y. Mely, Structure of the His44 → Ala Single Point Mutant of the Distal Finger Motif of HIV-1 Nucleocapsid Protein: A Combined NMR, Molecular Dynamics Simulation, and Fluorescence Study†Biochemistry 43 (2004) 7687-7697.
T. Shimizu, T. Hiyama, M. Ikeuchi, Y. Inoue, Nucleotide sequence of a metallothionein gene of the thermophilic cyanobacterium Synechococcus vulcanus, Plant Molecular Biology 20 (1992) 565-567.
K.O. Stetter, Extremophiles and their adaptation to hot environments, FEBS Letters 452 (1999) 22-25.
T.D. Brock, Micro-organisms adapted to High Temperatures, Nature 214 (1967) 882-885.
D.M. Ward, M.J. Ferris, S.C. Nold, M.M. Bateson, A Natural View of Microbial Biodiversity within Hot Spring Cyanobacterial Mat Communities, Microbiol. Mol. Biol. Rev. 62 (1998) 1353-1370.
S.R. Miller, R.W. Castenholz, Evolution of Thermotolerance in Hot Spring Cyanobacteria of the Genus Synechococcus, Appl. Environ. Microbiol. 66 (2000) 4222-4229.
U. Nübel, F. Garcia-Pichel, M. Kühl, G. Muyzer, Spatial scale and the diversity of benthic cyanobacteria and diatoms in a salina, Hydrobiologia 401 (1999) 199-206.
A. Hiraishi, T. Umezawa, H. Yamamoto, K. Kato, Y. Maki, Changes in Quinone Profiles of Hot Spring Microbial Mats with a Thermal Gradient, Appl. Environ. Microbiol. 65 (1999) 198-205.
Temperature on Species Composition and Community Structure of Hot Spring Microbial Mats, Appl. Environ. Microbiol. 66 (2000) 2835-2841.
J. Felsenstein, Phylogenies and the Comparative Method, The American Naturalist 125 (1985) 1.
R.W. Castenholz, Thermophilic blue-green algae and the thermal environment, Microbiol. Mol. Biol. Rev. 33 (1969) 476-504.
L.K. Larisa, H. Ibolya, V. Làszlo, A.L. Dmitry, Temperature-induced specific lipid desaturation in the thermophilic cyanobacterium Synechococcus vulcanus, FEMS Microbiology Letters 175 (1999) 179-183.
S. Silver, T.K. Misra, Plasmid-Mediated Heavy Metal Resistances, Annual Review of Microbiology 42 (1988) 717-743.
R.W. Olafson, W.D. McCubbin, C.M. Kay, Primary- and secondary-structural analysis of a unique prokaryotic metallothionein from a Synechococcus sp. cyanobacterium, Biochem. J. 251 (1988) 691-690.
J.D. Otvos, R.W. Olafson, I.M. Armitage, Structure of an Invertebrate Metallothionein from Scylla-Serrata, J. Biol. Chem. 257 (1982) 2427-2431.
S. Belkin, Microbial whole-cell sensing systems of environmental pollutants, Current Opinion in Microbiology 6 (2003) 206-212.
I. Bontidean, J.R. Lloyd, J.L. Hobman, J.R. Wilson, E. Csöregi, B. Mattiasson, N.L. Brown, Bacterial metal-resistance proteins and their use in biosensors for the detection of bioavailable heavy metals, Journal of Inorganic Biochemistry 79 (2000) 225-229.
P. Corbisier, D. van der Lelie, B. Borremans, A. Provoost, V. de Lorenzo, N.L. Brown, J.R. Lloyd, J.L. Hobman, E. Csöregi, G. Johansson, B. Mattiasson, Whole cell- and protein-based biosensors for the detection of bioavailable heavy metals in environmental samples, Analytica Chimica Acta 387 (1999) 235-244.
M. Valls, V. de Lorenzo, Exploiting the genetic and biochemical capacities of bacteria for the remediation of heavy metal pollution, FEMS Microbiology Reviews 26 (2002) 327-338.
D.P. Higham, P.J. Sadler, M.D. Scawen, Cadmium-Resistant Pseudomonas putida Synthesizes Novel Cadmium Proteins, Science 225 (1984) 1043-1046.
M. Mejáre, L. Bülow, Metal-binding proteins and peptides in bioremediation and phytoremediation of heavy metals, Trends in Biotechnology 19 (2001) 67-73.
M. Valls, V. de Lorenzo, R. Gonzàlez-Duarte, S. Atrian, Engineering outer-membrane proteins in Pseudomonas putida for enhanced heavy-metal bioadsorption, Journal of Inorganic Biochemistry 79 (2000) 219-223.
M. Valls, R. González-Duarte, S. Atrian, V. De Lorenzo, Bioaccumulation of heavy metals with protein fusions of metallothionein to bacteriol OMPs, Biochimie 80 (1998) 855-861.
S. Koji, Y. Yuusuke, H. Naoaki, Construction of a marine cyanobacterial strain with increased heavy metal ion tolerance by introducing exogenic metallothionein gene, Journal of Marine Biotechnology 6 (1998) 174-177.
Additional References cited in the text
G.J. Brewer, Recognition, Diagnosis, and Management of Wilson’s Disease, Proceedings of the Society for Experimental Biology and Medicine 223 (2000) 39-46.
C. Tohyama, Z.A Shaikh, K. Nogawa, E. Kobayashi, R. Honda, Urinary metallothionein as a new index of renal dysfunction in “Itai-Itai” disease patients and other Japanese women environmentally exposed to cadmium, Archives of Toxicology 50 (1982) 159-166.
H.S. Houn, M.L. Brantly, C. Oliver, M. Adamson, S.G. Kaler, W.A. Gahl, Metallothionein Synthesis and Degradation in Indian Childhood Cirrhosis Fibroblasts Pediatric Research 35 (1994)197-204.
M.T. Pellecchia, C. Criscuolo, K. Longo, G. Campanella, A. Filla, P. Barone Clinical Presentation and Treatment of Wilson’s Disease: A Single-Centre Experience, European Neurology 50 (2003) 48-52.
Y. Uchida Growth-Inhibitory Factor, Metallothionein-Like Protein, and Neurodegenerative Diseases, Biological Signals 3 (1994) 211-215.
S. Tsuji, H. Kobayashi, Y. Uchida, Y. Ihara, T. Miyatake, Molecular cloning of human factor growth inhibitory cDNA and its down-regulation in Alzheimer’s disease, The EMBO Journal. 11 (1992) 4843- 4850.
J.R. Hawse, J.R. Cumming, B. Oppermann, N.L. Sheets, V.N. Reddy, M. Kantorow, Activation of Metallothioneins and α-Crystallin/sHSPs in Human Lens Epithelial Cells by Specific Metals and the Metal Content of Aging Clear Human Lenses, Investigative Ophthalmology & Visual Science 44 (2003) 672-679.
K. Balamurugan, W. Schaffner, Regulation of Metallothionein Gene Expression, Metal Ions in Life Sciences 5 (2009) 31–49.
L. Chen, L. Lei, T. Jin, M. Nordberg, G.F. Nordberg, Plasma Metallothionein Antibody, Urinary Cadmium, and Renal Dysfunction in a Chinese Type2 Diabetic Population, Diabetes Care 29 (2006) 2682-2687.
C. Diorio, J. Cai, J. Marmor, R. Shinder, M.S. DuBow, An Escherichia coli Chromosomal ars Operon Homolog Is Functional in Arsenic Detoxification and Is Conserved in Gram-Negative Bacteria, Journal of Bacteriology 177 (1995) 2050–2056.
B. Gold, H. Deng, R. Bryk, D. Vargas, D. Eliezer, J. Roberts, X. Jiang, C. Nathan, Identification of a Copper-Binding Metallothionein in Pathogenic Mycobacteria, Nature Chemical Biology 4 (2008) 609-616.
K.G. Daniel, R.H. Harbach, W.C. Guida, Q.P. Dou, Copper Storage Diseases: Menkes, Wilson’s, and Cancer, Frontiers in Bioscience 9 (2004) 2652-2662.
H.V. Perales-Vela, J.M. Pena-Castro, R.O. Canizares-Villanueva, Heavy metal detoxification in eukaryotic microalgae, Chemosphere 64 (2006) 1-10.