Abstract
An increase in the worldwide consumption of fuel has created anxieties regarding diminishing reserves of fossil fuels. Additionally, the use of fossil fuels is linked to adverse environmental consequences. As a result, there is an increased interest in the use of biofuels, which are sustainable fuel sources with minimal environmental concerns. Microbial systems are commonly being used for the production of biofuels. OleTJE is a unique variant of cytochrome P450, a special group of enzymes that catalyses the formation of terminal alkenes from fatty acids.
However, this process leads to the formation of undesired α- and β-hydroxylation by-products, which lower the yield of terminal olefins. The purpose of this project is to use quantum mechanics (QM) and molecular mechanics (MM) to engineer OleTJE to overproduce terminal olefins by eliminating these hydroxylation by-products. A literature review was conducted to understand the reaction mechanisms of OleTJE and identify pathway points for manipulation.
It was noted that successful blocking of α- and β-hydroxylation reactions would require changing the active site of the enzyme, temperature modifications, and experimenting with different OleTJE enzyme systems. Starting samples will be obtained from the protein databank in pdb format and manipulated to include the required components before being subjected to computational manipulations. It is expected that the successful implementation of this project will have positive personal, academic, industrial, and economic impacts.
Introduction
A rapid increase in global fuel consumption has generated numerous concerns regarding dwindling fuel reserves. This problem is exacerbated by volatile geopolitical factors, the inconsistent cost of fossil fuel, and increasing apprehensions about national energy security (Covert et al., 2016). Furthermore, the use of fossil fuel causes grave environmental concerns due to the emission of greenhouse gases (Nicoletti et al., 2015).
As a result, there is an urgent need to develop sustainable fossil fuel substitutes. Biofuels generated from biological resources exemplify a promising substitute for fossil fuels because they are renewable and do not have adverse environmental consequences. The most common biofuels in the current market are bioethanol and biodiesel. Nevertheless, the best biofuels are bio-hydrocarbons, particularly intermediate to long-chain fatty alkanes or alkenes (Liu et al. 2014).
Other uses of aliphatic hydrocarbons include polymers, detergents, and lubricants. Preference is given to biofuels of this type because of their close resemblance to petroleum-based fuels in terms of a chemical structure and physical attributes. Therefore, recent research efforts have focused on biosynthetic pathways from different microorganisms and their potential in the production of aliphatic hydrocarbons (Liao et al., 2016; Zargar et al., 2017). One benefit of using microorganisms to synthesize fatty alkanes or alkenes is that microbial systems are amenable to scalability, which makes it possible to develop cost-effective systems.
As with most biological processes, the production of biofuels in microbial systems involves biological reactions that require enzymes. Enzymes are biological catalysts that hasten the rate of biochemical reactions. Living organisms produce enzymes naturally to facilitate the conversion of substrates to valuable products that serve various biological functions. This conversion involves the binding of the enzyme to the substrate and an intricate series of other reactions.
Naturally occurring enzymes demonstrate high levels of efficiency in their various roles at biological temperatures and pressures by reducing the rate of reactions substantially. However, experimental conditions differ from physiological ones, which interfere with the catalysis of the enzyme outside biological systems. Consequently, rigorous research is ongoing to elucidate the properties and machinery of biological enzymes to facilitate the exploitation of biocatalysts in the chemical industry.
However, such endeavours have been thwarted by the nature of enzyme-catalysed reactions, which are too fast for investigators to obtain accurate details that would be beneficial for comprehending the machinery and reaction pathways. Due to this limitation, novel computational techniques have been established to create accurate models with the capacity to predict the reactions of biocatalysts. Hence it is now possible to examine large biochemical systems.
Studies on the production of biofuel in microbial systems have revealed the involvement of a special group of enzymes known as cytochrome P450 alongside heme enzymes. Cytochrome P450 is an important group whose main role is to metabolise drugs in the human body (Liu et al. 2014). This group of enzymes is ubiquitous in living organisms, including plants, fungi, bacteria, and eukaryotic organisms. Due to their involvement in different biological functions, different subclasses of cytochrome P450 have been characterized.
A heme enzyme reacts with fatty acid substrates in a reaction that involves a decarboxylation step mediated by a cytochrome P450 peroxygenase. This scheme has the capacity to synthesise biofuel (Grant et al., 2015). However, it liberates a number of by-products that must be removed from the desired product before it can be used. At present, most investigative efforts report 50% decarboxylation and 50% hydroxylation (Faponle et al., 2016). Therefore there is a need to engineer the protein to enhance the yield of biofuel.
The use of quantum mechanics or molecular mechanics (QM/MM) techniques have made it possible to characterize biochemical systems. Quantum-mechanical (QM) approaches fall into ab initio methods that entail the computation of a system devoid of prior knowledge (de Visser et al. 2014). Overall QM methods work by unravelling the Schrödinger equation. Even though QM methods make it possible to achieve accurate computation of systems, their requirements increase depending on the number of atoms or electrons in the system. For this reason, they are unsuitable for investigations involving enzymatic systems (de Visser 2009).
On the other hand, substitute methods such as molecular mechanics (MM) that can handle systems with a higher number of atoms are incapable of demonstrating bond-breaking or formation processes. As a result, novel techniques that blend the rapidity of MM with the precision of QM have been developed, leading to computational procedures that can explain enzymatic systems. One such method is based on the density functional theory (DFT) alongside high computational power, which has led to increased interest in theoretical chemistry processes.
DFT computations are often done to back experimental biochemical and inorganic investigations, aid in data interpretation, and direct new experiments. This proposal seeks to conduct a thorough literature review on the use of the OleT enzyme in biofuel production, determine existing gaps, and propose suitable methods to fill these gaps by engineering the enzyme to improve its yield.
Project Objectives
The aim of this project is to engineer the OleT enzyme to improve its biofuel yield.
Proposal objectives include:
- It is conducting a detailed literature review to understand the mechanisms that lead to the formation of products and by-products.
- It is using computational methods to design and develop mutants that give a higher yield of biofuel products.
- We were discussing the expected personal, academic, industrial, societal, and economic benefits of this project.
Background
Production of Biofuels
So far, four bacterial biosynthetic pathways that lead to the transformation of free fatty acids or fatty acid thioesters into bio-hydrocarbons have been classified. The first pathway is a cyanobacterial pathway comprising two enzymes: an acyl-acyl carrier protein (acyl-ACP) reductase and an aldehyde decarbonylase. These two enzymes change fatty acyl-ACPs to alkanes (Liu et al. 2014).
A second pathway is a group of three genes in Micrococcus luteus, which produces alkenes with internal double bonds via the head-to-head joining of two fatty acyl-coenzyme A (acyl-CoA) molecules (Chen et al. 2015). The third reaction path entails a distinctive P450 decarboxylase OleTJE, which was isolated from Jeotgalicoccus sp. ATCC 8456. This enzyme catalyses the direct decarboxylation of long-chain fatty acids to produce α-olefins in the presence of H2O2 (Liu et al., 2014). The last reaction involves a type I polyketide synthase isolated from Synechococcus sp. PCC 7002 (Lin et al., 2015).
This enzyme converts fatty acyl-ACPs into α-olefins through five successive steps. The one-step decarboxylation of fatty acids by OleTJE P450 is the simplest approach of all these methods. In addition, this method makes use of free fatty acids as substrates in the place of fatty acid thioesters, which is considered beneficial for metabolic engineering due to the abundance of free fatty acids (Faponle et al., 2016). Moreover, free fatty acids are amenable to manipulation in E. coli, which is one of the most advanced microbial cell industrial units (Wang & Zhu 2018). Hence, the P450 fatty acid decarboxylative mechanism holds immense potential for engineering into a biological α-alkene-producing scheme.
Cytochrome P450
Cytochrome P450 is a superfamily of heme-containing external monooxygenases (Faponle et al., 2016). The name of this group of enzymes stems from their classification as hemoproteins (they contain heme as a cofactor) and demonstration of distinct spectral characteristics. P denotes pigment, whereas the term 450 indicates the rare absorption peak of the reduced CO-bound compound at 450 nm.
Cytochrome P450s are crucial drug metabolising enzymes in the human body, primarily in phase 1 reactions (Faponle et al. 2016). They are also found in other living organisms, including eukaryotic animals, plants, fungi, and bacteria. This group of enzymes is very diverse, with over 20,000 distinct sequences identified as of January 2015. In humans, P450s are mainly found in the liver, where they catalyse the breakdown of xenobiotics (Ji et al. 2015).
However, they are also useful in the degradation of vitamin D, biosynthesis of hormones, metabolism of alkaloids, steroids, eicosanoids, and fatty acids, the formation of bile acid, as well as the instigation of procarcinogens, which demonstrates their multiple roles in the body and their flexibility in the activation of substrates. In plants, cytochrome P450 facilitates the metabolism of herbicides and secondary metabolites (Zhai et al., 2014).
The substrate-binding compartment of P450 differs in shape and size in various isozymes. The functional group of P450 enzymes consists of an iron (IV)–oxo heme cation radical moiety known as Compound I (Faponle et al., 2016). However, experimental investigation of compound I am complicated by its fleeting nature, which limits the availability of details about its catalytic machinery. A widespread reaction mechanism implemented by the P450 enzymes is aliphatic hydroxylation of substrates, which has been demonstrated empirically as a stepwise progression that forms alcohol product compounds.
This mechanism has been validated by computational modelling, which revealed that the oxidation state comprises two spin states (lying close to each other) that produce two-state reactivity configurations, each having its own rate constant (de Visser et al. 2014).
The two-state reactivity prototype also forecasts that the lifespan of the radical intermediary could cause a split of the potential energy. Cytochromes P450 take part in catalytic reactions, for instance, dehalogenation, peroxidation, hydroxylation, sulfoxidation, S-dealkylation, desulfuration, and epoxidation, among many others (Zhao et al. 2014). The interaction of these enzymes with their corresponding electron donors is crucial because this specificity guarantees adequate catalysis and satisfactory reaction rates while shielding the system from bypass reactions.
The cytochrome P450 enzyme superfamily is among the most adaptable naturally occurring biocatalysts. They use oxygen to catalyse their reactions. Electron-moving proteins convey electrons to the enzyme to expedite oxygen instigation, whereas NADH or NADPH facilitate the hydroxylation of substrates. Most cytochrome P450 variants are bound by membranes and have different hydrophobic substrates on which they catalyse monooxygenation reactions.
Some of the substrates involved in these reactions include biological compounds such as steroids, fatty acids, and prostaglandins. On the other hand, extraneous compounds involved in monooxygenation reactions include organic solvents, ethanol, anaesthetics, and alkyl aryl hydrocarbon products. Monooxygenase reactions involve the use of one or more redox affiliate proteins to move two electrons from NADH or NADPH to the heme iron in the reactive centre for dioxygen initiation. Thereafter, one atom of oxygen is inserted into their substrates, which leads to the name P450 monooxygenases. A typical cytochrome P450 catalytic cycle is shown below in Figure 1.
However, a unique cytochrome P450 family, the CYP152 family, contains members that use H2O2 entirely as the only electron and oxygen donor. These members are known as P450 peroxygenases, and they include P450SPα, P450BSβ, and OleTJE (Matthews et al., 2017a). The peroxygenase activity of P450 enzymes is usually considered an advantage in practical settings because H2O2 is significantly cheaper than NADPH and redox proteins. These applications encompass the use of P450 enzymes as catalysts for biosynthetic reactions in living organisms and enzyme additives in laundry cleansing agents (de Visser 2009). Therefore, there is a renewed interest in exploiting the oxidative role of cytochromes P450 for industrial goals.
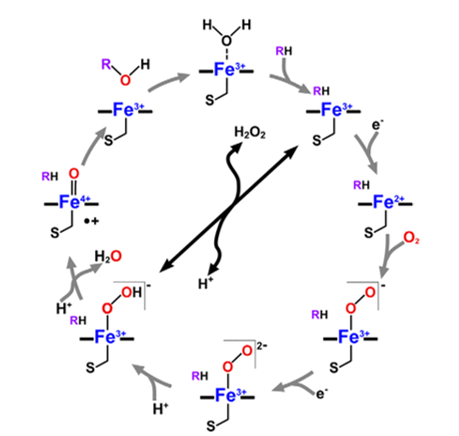
An important reaction mechanism in the chemistry of cytochrome P450 chemistry is the desaturation of aliphatic compounds to form olefins (Munro et al., 2018). Desaturation reactions are similar to aliphatic hydroxylation because both processes begin with the initial removal of a hydrogen atom (Ji et al., 2015). However, the reaction subsequently divides into two likely product routes: one presenting OH rebound to create alcohol products, and the other resulting in the removal of a second hydrogen atom to produce an olefin and water (Matthews et al. 2017a).
The first reaction pathway is referred to as aliphatic hydroxylation, whereas the second path is known as desaturation, which is of great interest in the production of biofuels. Even though comprehensive scientific investigations into the chemistry of P450 have been conducted for decades, many gaps exist in the understanding of its reaction mechanism.
Cytochrome P450 Peroxygenase OleTJE
Cytochromes P450 that use the peroxide shunt to catalyse the hydroxylation of long-chain fatty acids to produce alcohol is known as peroxygenases. Adding hydrogen peroxide or any other organic peroxide facilitates the production of compound 0 or I without requiring costly redox partners. The most important cytochrome P450 peroxygenases belong to the CYP152 family (Matthews et al. 2017b).
A notable peroxygenase is cytochrome P450 peroxygenase OleTJE, which was isolated from Jeotgalicoccus sp. ATCC 8456. This enzyme catalyses the extraordinary decarboxylation of long-chain fatty acids leading to the formation of α-alkenes. This process uses H2O2 as the sole electron and oxygen donor. Another reaction mechanism involves an enzymatic redox cascade using molecular oxygen and the recycling of NAPDH, as shown in Figure 2.
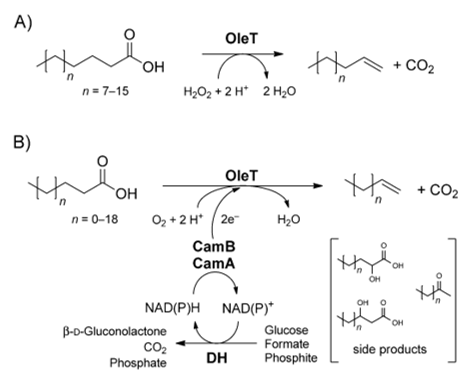
Nonetheless, it is not possible to manufacture cheap α-alkene biofuels on a large scale via the decarboxylase activity of OleTJE P450 while depending on the H2O2-dependent enzymatic system. This production is not feasible because using large quantities of peroxide is exorbitant. Additionally, high concentrations of H2O2 are detrimental to the wellbeing of biocatalysts, which become inactivated. As a result, the H2O2-independent action of OleTJE is ideal for the profitable microbial production of α-alkenes (Liu et al., 2014). Aliphatic α-alkenes are valuable because of their potential use as biofuel substitutes for fossil fuels, manufacture of lubricants, detergents, and polymers.
Therefore, studies on fatty acid decarboxylation by OleTJE are important and have the potential to result in the industrial manufacture of biogenic α-alkenes, which are renewable and environmentally friendly fuel sources.
H2O2-Independent Activity of OleTJE
H2O2-independent decarboxylation of fatty acids by OleTJE happens in vivo in the presence of NADPH and O2. Under these conditions, this enzyme efficiently decarboxylates long-chain fatty acids (with carbon atoms ranging from 12 to 20) to produce terminal olefins. This process requires OleTJE to work alongside a fused P450 reductase domain RhFRED from Rhodococcus sp., CamA, CamB, or a separate flavodoxin/flavodoxin reductase from E. coli.
The unearthing of the H2O2-free activity of OleTJE probes the monooxygenase-like machinery of the peroxygenase and guides future metabolic engineering efforts to enhance redox partners for efficient production of α-alkenes by OleTJE. However, the reaction liberates significant quantities of α- and β-hydroxylation by-products whose origin is not well understood. The independent peroxide activity of OleTJE is a promising approach to the production of biofuels because it safeguards the wellbeing of microbial systems.
α- and β-Hydroxylation by-products of OleTJE
QM/MM investigations on the bifurcation pathways have made it possible to identify and elucidate the generation of three by-products of OleTJE. These studies have also made it possible to demonstrate how the enzyme can be engineered for optimal desaturase activity. It is evident that the polarity and solvent availability of the substrate in the binding cleft weakens the OH-rebound path and allow a thermodynamically unfavourable decarboxylation reaction through kinetic means, as indicated in Figure 3.
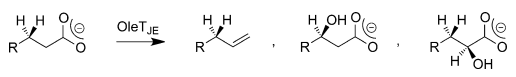
Substrate Specificity of OleTJE Systems in Peroxide-Independent Pathways
The substrate preference of various OleTJE systems provides valuable data to enhance the comprehension of the in vivo machinery of fatty acid decarboxylases. This information directs the metabolic engineering of fatty acid biosynthesis. Various straight-chain saturated fatty acids with even numbers of carbon atoms in the hydrocarbon chain have been used to determine the substrate specificity of different OleTJE systems. Liu et al. (2014) used long-chain fatty acids ranging from C8 to C20 to determine the substrate specificity of OleTJE and OleTJE-RhFRED.
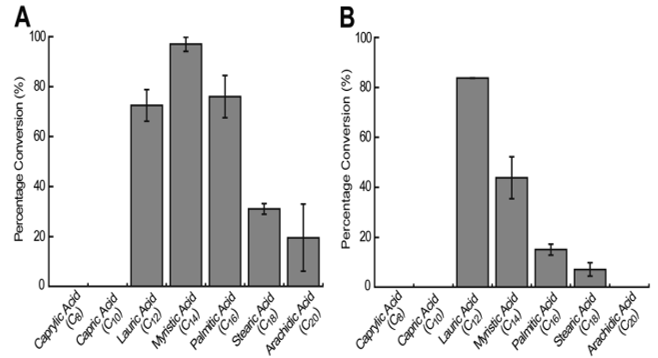
The percentage conversion of substrate into the matching α-alkene product was used to ascertain the substrate preference. It was noted that OleTJE had the highest preference for myristic acid (C14) for olefin production with a conversion ratio of 97%. The conversion of lauric acid (C12) and palmitic acid (C16) was below the ideal levels. OleTJE could only transform a small quantity of stearic acid (C18) and arachidic acid (C20) to alkenes (Figure 4). However, it could not catalyse the decarboxylation of capric acid (C10) or caprylic acid (C8).
Conversely, it was shown that OleTJE-RhFRED had a higher preference for fatty acids with relatively short chain lengths. This observation implied that the P450-reductase contact might produce a small structural change in the active site of OleTJE. The highest activity for OleTJE-RhFRED was shown against lauric acid at a conversion rate of 83.8%. However, no activity was observed for arachidic, capric, and caprylic acids (Li et al., 2014). Another notable observation was that the system consisting of OleTJE-RhFRED, NADPH, and O2 exhibited a lower activity than OleTJE and H2O2 when testing all fatty acids except lauric acid. This phenomenon points towards the likelihood of OleTJE evolving over time to enhance its peroxygenase activity.
On the other hand, Dennig et al. (2015) demonstrated the enzymatic oxidative decarboxylation of short-chain fatty acids ranging from C4 to C9 using OleT, O2 as the oxidant, and NAD(P)H as the electron donor to generate the equivalent terminal C3 to C8 alkenes. Product titres of up to 0.93 gL-1 and total turnover numbers exceeding 2000 were given off. Attaining these yields was made possible by developing a well-organized electron-transfer series using putidaredoxin CamAB alongside the utilisation of NAD(P)H without regard for formate, glucose, or phosphite (Dennig et al. 2015).
As much as CamAB could oxidize NAD(P)H without an appropriate electron acceptor, no H2O2 was detected following full oxidation of 1 mm NADH, which pointed towards a direct electron displacement from CamAB to OleT. This observation also eliminated the possibility of inactivation by hydrogen peroxide. For this reason, the nonexistence of catalase did not affect the throughput of the system. Surprisingly, fatty acids with short chains (less than 9 carbon atoms from C4 to C11) were decarboxylated, thereby leading to the production of 0.04 to 2.45 mm of terminal alkenes. The total yield was influenced by the prevailing temperature and the chain length of the substrate. The highest activity was recorded for a C18 fatty acid at room temperature, whereas C12 yielded the best outcome at 48°C (3.26 mm of 1-undecane).
On the whole, fatty acids ranging from C10 to C16 had high conversion rates at 48°C. This system enabled the production of propene from butyric acid in only one step, which refuted the former hypothesis that OleT was specific to long-chain fatty acids. It was assumed that long-chain fatty acids associated with the substrate channel, while smaller substrates were held in the binding compartment as evident in correlated enzymes, for example, CYPBSß.
The poor action of fatty acids with intermediate chain lengths (C10 and C11) was attributed to poor binding on the enzyme. Decarboxylation was the principal reaction, with approximately 86 to 99% selectivity in fatty acids C18 to C22. The formation of α- and β-hydroxylated products was influenced by chain length and reaction temperature. There was 62% α-/β-hydroxylation when fatty acids with 9 carbon atoms reacted at 48°C.
The observed harmony between CamAB and OleT was attributed to similarities in the coordination of the adjacent cysteine thiolate moieties in the ferrous states of OleT and P450Cam, which permit the attachment of oxygen to the heme iron. Most P450 enzymes of bacterial origin depend on class I electron-transfer schemes (Dennig et al. 2015), which accounts for the enhanced catalytic performance of OleT together with CamAB, as opposed to systems illustrated in the past, such as OleT–RhFRed and Fdr/Fdx. These findings indicate that metabolic engineering efforts directed at the peroxide independent activity of OleTJE should consider pairing this enzyme with CamAB to take advantage of different lengths of fatty acid substrates.
Product Distribution in OleTJE Catalysis
The branching and regioselectivity inclinations of enzymes are sometimes affected by environmental effects. Therefore, Faponle et al. (2016) used QM/MM to study the full OleTJE enzyme and investigate its bifurcation pathway and product distribution. The main objectives were to identify the source of P450 OleTJE product formation and predict the features that separate the decarboxylation from hydroxylation pathways. There were no significant changes in geometry, spin state collation, and comparative energies. Geometry studies showed that there were no substantial disparities between hydrogen-atom extraction impediments.
In all instances, the hydrogen atom was nearly equidistant from the donor and acceptor atoms, notwithstanding the type of the QM region studied. It was evident that hydrogen atom removal impediments by Cpd I from the Cα and Cβ loci of the substrate were close in energy on the doublet or quartet spin states. All four impediments fell within a range of 3.2 (BS1) or 4.0 (BS2) kilocalories per mole for Sn400 (Faponle et al., 2016).
The observed hydrogen atom extraction impediments corroborate empirical findings of a combination of reaction outcomes from Cα and Cβ hydrogen atom removal. The slight energy difference between the two pathways implies that the hydrogen atom removal reactions between Cα and Cβ are competitive. However, the ratio of the Cα and Cβ pathways as depicted by the comparative energies of the transitional state moieties 4,2TSHA,α and 4,2TSHA,b were strongly influenced by environmental factors, for example, the associations between hydrogen bonds (Faponle et al. 2016).
This experiment showed that hydrogen atom extraction was the rate-limiting stage in the reaction mechanism. Therefore, the authors approximated a large kinetic isotope effect and computed an estimate for the substitution of hydrogen atoms by deuterium (molecular mass of 11). Geometry shots were also taken to determine possible hydrogen atom removal from the Cγ locus of the substrate (Sn400). It was noted that the reaction energy exceeded the reactant complex by more than 30 kcal mol-1 hence this mechanism was overlooked (Faponle et al., 2016).
The system slowed down following the removal of hydrogen atoms to form a radical intermediate (IH,α or IH,β) with an iron(IV)–hydroxo group and a reactive substrate on either the Cα or Cβ locus of the substrate. The transitional species 4,2IH,α only reacted through radical rebound past the rebound transition state 4,2TSreb,α thereby leading to the formation of α-hydroxo arachidonic acid upshots (4,2 P OH,α).
The rebound barriers recorded during this investigation were larger and higher in energy than is the norm for P450 hydroxylation reactions. Under normal circumstances, the low-spin pathway does not have any impediments. A decarboxylation reaction cannot take place following the removal of Cα hydrogen; it requires the additional removal of a hydrogen atom from Cβ to Cα, which requires high energy.
The competitive pathways leading to decarboxylation and hydroxylation products (PD,β and POH,β, correspondingly) via transition states TSD,β and TSreb,β came from the radical intermediate 4,2IH,β, as estimated by snapshots Sn300 and Sn400. The high-spin pathway through 4 IH,β gave rebound, and decarboxylation hindrances that were almost similar in energy hence could be regarded as competitive.
On the other hand, the low-spin path produced quality decarboxylation products from 2TSH,β directly without stable radical intermediates. These findings indicated that the rebound barriers were bigger than conventionally found in ideal complexes, which was attributed to numerous hydrogen-bonding connections to the iron–hydroxo group. These restrictions increased the obstacles in the rotation and rebound pathways. In addition, the hydroxylation process was dominant following Cα-H removal, while decarboxylation was favoured following the extraction of the Cβ-H atom on the doublet spin state.
On the other hand, the surface of the quartet spin state was expected to liberate a mixture of products. This observation implies that the reaction should be modified to promote Cβ-H extraction and impede Cα-H removal to promote decarboxylation. The elevated rebound barrier was linked to hydrogen-bonding interactions because of the water channel in P450 OleTJE in addition to the polarity of the nearby carboxylate-Arg245 salt bridge (Faponle et al. 2016). The importance of water channels in P450 OleTJE was also demonstrated.
Bifurcation Pathways of OleTJE
Faponle et al. (2016) used a valence-bond (VB) diagram to describe the bifurcation pathways of P450 OleTJE (Figure 5). In this illustration, the wave function of the reactant (YI) joins an energised state in the geometry of the yields, which is Yreb* for the alcohol upshots and YD* for the outcomes of decarboxylation. In the same way, the wave function of the product joins the energised excited state in the reactant geometry (4,2IH,β).
The magnitude of the reaction impediment depends on the excitation energy required to transform the reactant to the product in the reactant geometry. Consequently, the bifurcation machinery is determined by the comparative magnitude of the excitation energies Greb or Gdecarb (Fapone et al., 2016). Therefore, the microelectronic disparities between the wave functions in the ground and excited states influence the barrier impediments and rate constants for the rebound and decarboxylation reactions.
An assessment of the VB structures in the geometry of 4,2IH,β with emphasis on the ground and excited states provides an understanding of the strictures that control the bifurcation variables.
For OH rebound and the development of hydroxylation outcomes, the πyz/π*yz set of orbitals was divided into two atomic orbitals, 3d yz,Fe and 2py,O. This process expended an estimated Eπ/π*yz in energy from the system, which was estimated at 37.2 kcal mol-1. The 2py,O orbital coupled with ΦSub, a substrate radical, to create the C-O bond orbital (σC-O) in a process that consumed a bond severance energy BDECO of 87.2 kcal mol-1.
On the other hand, since the 2py,O had a double occupancy in 4,2IH,β, it was imperative to upgrade one electron to a free low-lying orbital. The two available orbitals were 3dxz in the doublet spin state and the σ*z2 orbital in the foursome spin state, Eexc. This excitation energy was approximated at 75.3 kcal mol-1. Thus the value of Greb was predicted as 25.3 kcal mol-1. As a rule, the height of the barricade is usually 33.33% of the promotion gap, which implied that a rebound barrier of approximately 8 kcal mol-1 was applicable.
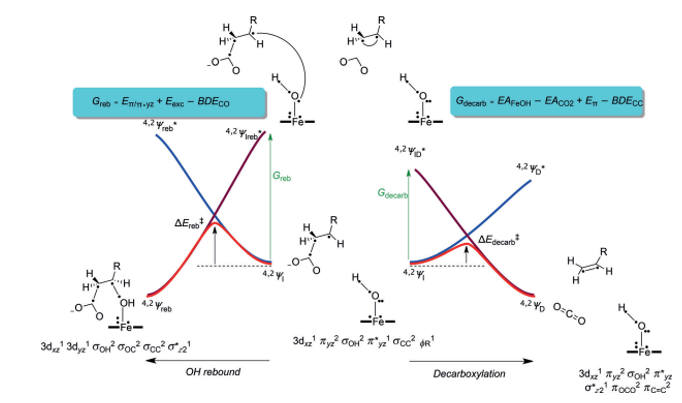
However, a different scenario was witnessed in the decarboxylation reaction, where two processes occurred simultaneously. The C-Cα bond of the substrate disintegrated (BDECC) while the leaving CO2– group was oxidised to CO2. This second process denoted the electron affinity of CO2 (EACO2). As a result, the electron duo in the C-Cα orbital (σCCα) was split, after which the electron on Cα coupled up with the reactive group on Cβ to create a π bond (π CαCβ) possessing energy Eπ.
The additional electron was taken up by the iron–hydroxo compound, which was reduced by an amount of energy that corresponded to its electron affinity (EAFeOH). The electron affinity of the iron complex was estimated at 56.5 kcal mol-1, whereas the strength of the C-C bond in the substrate, BDECC, was found to be 7.8 kcal mol-1 (Faponle et al., 2016). Conversely, the singlet to triplet energy gap was used to compute the energy of the π bond, which yielded a value of 73.7 kcal mol-1. A promotion gap for decarboxylation (Gdecarb) of 3.4 kcal mol-1 was computed using the standard electron affinity of CO2 (‑0.60 eV).
The findings pointed towards a minor reaction barrier that was estimated to be less than 1 kcal mol-1. Figure 6 indicates the calculated bifurcation impediments of different radical intermediates.
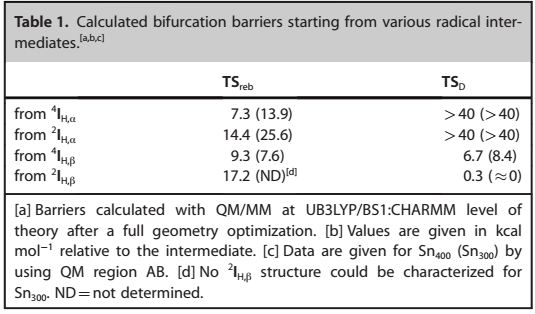
Factors Affecting Decarboxylation Reactions of OleTJE
Before designing an enzyme to overproduce a given product, it is important to determine the factors that promote the reaction pathway that leads to the desired product. Investigations show that the decarboxylation pathway of OleTJE is mainly influenced by substrate factors (Xu et al., 2017). These aspects include the intensity of the C-Cα bond, the energy required to create the π bond between Cα-Cβ, and the electron affinity of CO2 (Faponle et al. 2016; Fang et al. 2017).
The impact of the oxidant on the decarboxylation reaction only happens via the electron affinity of the iron (IV)–hydroxo moiety. Consequently, enhancing the regioselectivity of decarboxylation as opposed to hydroxylation needs to concentrate on weakening the hydroxylation path by engineering the substrate-binding compartment or using alternate substrates. The computations presented by Faponle et al. (2016) show that the substrate-binding compartment of P450 OleTJE is highly polar and has numerous hydrogen-bonding interactions, which are crucial to the disruption of the radical rebound reactions that lead to the formation of α- and β-hydroxo fatty acids.
In addition, electron-extracting moieties attached to the Cα or Cβ locus would elevate the energy required to form the C-O bond, which would increase the radical rebound barriers. In contrast, the decarboxylation reactions would be boosted further by using substrates with an alkene bond between Cγ and Cδ. Such substrates would promote the π conjugation with the Cα-Cβ π bond, thereby leading to increased Eπ energy and stabilisation of the decarboxylation process.
This literature review has provided useful insights that will guide the engineering of OleTJE to enhance its productivity of biofuels. The following insights have revealed that the enhancement of olefin product distributions will require site-directed mutations that impede α- and β-hydroxylation as well as altering the substrate-binding compartment to limit hydroxylation rebound reactions (Li et al. 2015). It is also evident that the choice of substrate, reaction temperature, and OleTJE systems or combination of systems (Dennig et al. 2015) is critical in obtaining optimal terminal olefin yields from fatty acids. These pointers will inform the computational methods in the following section.
Methodology
Computational modelling of biosystems provides researchers with a basis to establish their experiments because it eliminates redundant variables that would otherwise be a waste of resources. Quantum mechanics/molecular mechanics (QM/MM) techniques have become popular over the years due to their capacity to provide data regarding the starting point of products or by-products through calculated reaction machinery and energy settings (de Visser et al. 2014).
As a result, it is possible to collect data regarding factors that influence enzyme reactions, rates, and specificities by analysing the different states of the molecules involved in the reactions. On the other hand, cluster replicas such as the Density Functional Theory (DFT) enable voluminous computations in a short span by analysing precise features of a single protein (de Visser 2009; Melander et al. 2016).
Even though cluster models shed light on the intrinsic activity of enzymes’ active sites, problems result from key bonding patterns and lengths. This section of the proposal provides a detailed procedure using general QM/MM optimization protocols to promote the decarboxylation mechanisms of cytochrome P450 OleTJE in the production of terminal olefins. The proposed elements of improvement, as highlighted in the literature review, will be incorporated into the procedure to determine the most effective manipulations that yield favourable outcomes.
QM/MM Methodology
QM/MM methods start by finding heavy-atom coordinates of the protein (enzyme) of interest from X-ray crystallographers in the literature or the protein databank library. These coordinates will be obtained in a pdb format, which implies that the structure will require editing and additional modifications prior to running simulations. Examples of relevant modifications include the addition of hydrogen atoms, missing chains, amino acid residues, and substrates. Crystal additive buffers should also be removed. The structure of the active site can also be altered, after which the system can be solvated.
These procedures permit the stabilization of the system to promote the proper folding of proteins. A molecular dynamics (MD) simulation is then executed to produce snapshots that are necessary to implement the QM/MM computations, which split up the biochemical system into inner and outer cores. The computations of the inner core are usually done using ab initio or DFT techniques that have high levels of accuracy (Lewars 2016). On the other hand, outer core calculations are effected using MM approaches. These procedures have been authenticated with experimental data many times by Quesne et al. (2016).
Selection of Starting Structure in pdb Format
It is imperative to observe high levels of accuracy and caution when choosing the starting crystal structure from the protein databank. At the same time, it is difficult to find the ideal structure directly from the protein databank, which implies that modifications are inevitable. However, it is necessary to find the most comprehensive structure to reduce the time required for modifications. Overall, pdb files indicate whether atoms and residues are absent.
Thus it is the user’s responsibility to ascertain whether the missing attributes are protein side-chains, loop areas, substrates, binding proteins, co-factors, or reducing partners and select the file with the closest semblance to the preferred structure. It is also necessary to select high-resolution pdb files (minimum resolution of 2 Å) to ensure accurate identification of atoms and their loci in the protein. In addition, the investigator will prefer files with Rfree values of 0.3 or lower to ensure that reliable structures are chosen.
Modifying the Starting Structure
The most informative data on enzymatic reactions are obtained from QM/MM computations of fast reactions. However, the pace of these reactions and their corresponding short-lived intermediates complicates finding their crystal coordinates. Therefore, a previous structure in a resting state will be taken and modified appropriately. Starting structure modifications that will be relevant in this process will be informed by the findings and recommendations of previous studies.
The modifications will involve substrate modification and alteration of the enzyme’s active site. Faponle et al. (2016) proposed that enhancing the regioselectivity of decarboxylation as opposed to hydroxylation reactions should focus on weakening the hydroxylation path by engineering the substrate-binding compartment or using alternate substrates. The computations presented by Faponle et al. (2016) show that the substrate-binding compartment of P450 OleTJE is highly polar and has numerous hydrogen-bonding interactions, which are crucial to the disruption of the radical rebound reactions leading to the formation of α- and β-hydroxo fatty acids.
Therefore, it is hypothesized that a further increase in the polarity of P450 OleTJE’s active site will impede the reactions leading to hydroxylation products. Another potential modification is attaching electron-extracting moieties to the Cα or Cβ locus to elevate the energy required to form the C-O bond and increase the radical rebound barriers.
Substrate Docking Protocols
The binding of a substrate to an enzyme triggers a conformational modification in the three-dimensional organization of the resultant enzyme-substrate complex. However, the pace of this process is too fast to permit the crystallisation of the analogous structure. Molecular and computational docking procedures have been established to forecast the orientations and loci of binding substrates.
An online server, SwissDock, which is available at www.swissdock.ch, will be used for this step. Docking procedures take advantage of basic data regarding the substrate and protein surface, including the types and configurations of atoms, atomic alterations, and polarity. These data are compared to find the most suitable lock-and-key pairs. The enzyme and substrates will be tested in each likely orientation, followed by the computation of energies for each alignment using MM methods. The probable enzyme-substrate complex will be identified by selecting the arrangement with the least energy. Software packages such as Autodock will be useful in this regard (Quesne et al., 2016).
The Addition of Hydrogen Atoms and Protein Groups to the Structure
Hydrogen atoms will be added to the protein structure in the pdb file following the execution of geometrical optimizations. A number of online programs and packages will be beneficial in this process to enhance the detectability of the right positions of hydrogen atoms. The addition of hydrogen atoms is determined by the oxidation states, pKa systems, and hybridisation of the atoms.
Selection of Molecular Modelling Replicas
The right field force needs to be used to minimize the energy of bonds before conducting QM/MMM computations. Energy minimisation promotes the attainment of correct bond lengths and hybridisation structures. Several field forces are available. However, since this project involves an enzyme, a biochemical field force such as GROMOS, CHARMM, or AMBER will be ideal for the process. The force field calculations will make use of the following equations.
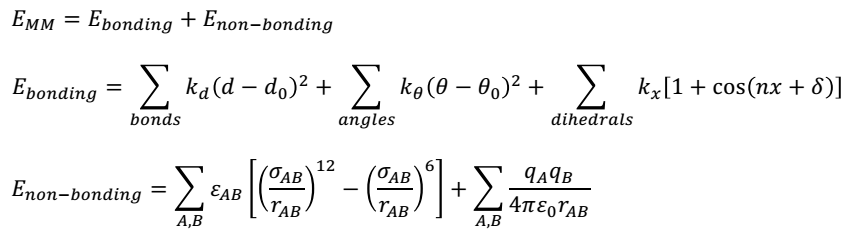
The first equation shows that MM energy (EMM) is the sum of bonded and non-bonded interactions. In the above equations, d denotes the bond length, ϴ shows the angles of valence bonding, and x stands for torsional dihedral angles. The dihedral attribute is a cosine function that takes into account the torsional multiplicity (n) and phase (δ) with respect to a bonding constant (kx). The non-bonding energy between two atoms A and B is spread between two interactions: a Lennard-Jones expression and a Coulomb interaction, as shown in the third equation.
The Lennard-Jones expression includes the rAB term that specifies the interatomic distance, whereas ϵAB and σAB describe the depth, breadth, and location of the dormant well. On the other hand, the (qAqB) term represents the effective atomic charge of the two atoms A and B, whereas ϵ0 describes permittivity in a vacuum. The value of qAqB is directly proportional to the inverse distance between the two atoms and ϵ0.
Solvation of the Protein Structure
Crystallised protein structures in pdb format lack some of the water molecules that should ordinarily be available. This lack arises from the fact that most crystallisation approaches require the elimination of water from protein suspensions (Jindal & Warshel 2016). Consequently, it will be necessary to add water to the initial starting structure to re-solvate the starting structure to simulate a lifelike enzyme model at room temperature. Re-solvation also improves the in vivo and in vitro portrayal of polarity. The major shortcoming of available resolving methods is that it is difficult to get water molecules to the inner regions of the protein. In addition, water molecules have the tendency to leave pits in the boundary of the protein, which interferes with the calculations (Jindal & Warshel 2016).
Therefore, it is necessary to re-solvate the system several times until no more water molecules can be added to the protein structure. Molecular, dynamic equilibration methods will be used to transfer water molecules into the pits. One method requires conducting a long MD simulation while exerting pressure from the perimeters of the system to compel the conveyance of water into the centre of the protein. The loci of the added water molecules will be checked following solvation to ensure that no hydrogen bonds have formed using atoms required in the reaction pathway.
Heating, Equilibrating and Conducting Molecular Dynamics Simulation on the Solvated Structure
Heating a chemical system is necessary to elevate its temperature, equilibrate the complex, and conduct MD simulations. The temperature of the system will be raised gradually at small intervals ranging from 0°K to 298°K. This process uses the applied force field to minimise the hydrogen atoms and solvent molecules. When the desired temperatures are achieved, the protein backbone becomes flexible, which equilibrates the system. At this point, MD simulations will be run for 500 to 1000 picoseconds to generate snapshots of the protein structure, reaction intermediates, and products for the required QM/MM computations. Different temperatures will be investigated to determine the impact of temperature on the decarboxylation mechanism of OleTJE.
Selection of Snapshots from the MD Simulation
It will be necessary to select a number of snapshots from the MD simulation because each snapshot will provide different calculations with respect to the reaction rate, barrier height, and product formation. In addition, changes in the patterns of hydrogen bonds within a single water molecule are likely to alter the total energy by approximately 4 kcal mol-1 (Hayashi et al., 2017). Therefore, several snapshots will permit the evaluation of varying conformations of the protein.
A number of MD snapshots will be taken from various points alongside the MD path graph. Each recreation is considered a starting point for the development of reaction profiles. Additionally, snapshots from different protein milieus will have varying parameters such as local minima of comparative energies, spin-state gaps, and order. Therefore, the average of the findings will be obtained when evaluating reaction schemes from more than one snapshot.
Selection of the QM Region, QM Methods, and Basis Set
The energetics of a reaction path and the electronic configuration of the active enzyme is influenced by the choice and size of the QM region. A small QM region can lead to incorrect descriptions of the polarisation of the system. Conversely, extremely large QM regions can lead to excessive polarisation. These outcomes are attributed to disparities in bond lengths when comparing QM regions of varying sizes. Therefore, it will be necessary to scrutinise numerous QM regions with diverse sizes to ascertain consistencies and select the appropriate regions.
Electrostatic and Mechanical Embedding Methods
When computing energies in the QM region, it will be necessary to consider the charges in the MM zone. Consequently, electrostatic or mechanical-embedding systems will be helpful. The mechanical-embedding procedures determine van der Waals forces and electrostatic influences of the MM atoms in the QM zone (Jindal & Warshel 2016). Geometric strictures such as the expanse of van der Waals interactions will be needed to separate the QM and MM energies. The QM/MM expressions will subsequently be incorporated directly into the force field estimations.
Calculation of QM/MM Energies
QM/MM energies will be calculated by completing geometrical optimisations on the QM and MM zones. Two methods of optimisation are possible: adiabatic and diabatic (Duarte et al., 2015). The adiabatic geometry optimisation approach entails relaxing the macro element exchanges inside the QM core as well as the micro-component interactions brought about by the neighbouring MM environment. In contrast, the diabatic style encompasses freezing the QM core region during the optimisation of the MM environment and vice versa. However, the total of the QM and MM energies do not account for the entire QM/MM energy of a system because linker atoms intermingle in the boundary of the two regions, thus producing energies that should be considered in the computations. These coupling terms are described by the following equation.
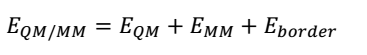
QM/MM computations can be achieved by assimilating an MM into a QM software package, for example, the ONIOM program, which is part of the Gaussian software package. These calculations can be executed using additive and subtractive procedures, as shown in the following equations.
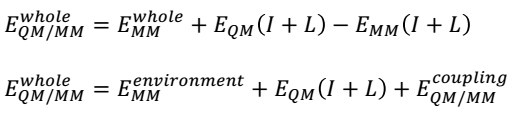
The subtractive approach, which will be used in this project, involves using linker atoms (L) to conceal the inner subsystems. The EQM (I + L) term symbolises the sum of the QM region (I) and linker atoms (L) at the QM level. Conversely, the EMM (I + L) term denotes a similar calculation at the MM level. However, it is important to subtract the MM energy of the inner subsystem from the energy of the entire system (E whole MM) to preclude double tallying of energies. The main benefit of the subtractive method is that it does not need a QM/MM joining term (Duarte et al., 2015).
Resources
Computational resources for this project, for example, software such as Autodock, CHARMM, DL_POLY, and TURBOMOLE, and hardware required to implement the computations, are already licensed by the University of Manchester, thus no additional costs will be incurred. Workspace is also available at the School of Computer Science at the University of Manchester. The primary researcher is a master’s student at the University of Manchester. Therefore, he will not require any compensation for the services offered. The primary supervisor will be remunerated by the University of Manchester.
Planning
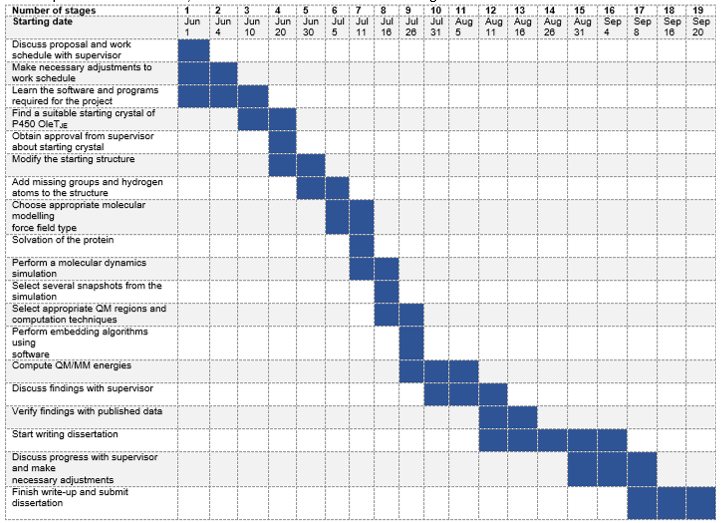
Figure 7 provides a rough guide to schedule until completion of the project, which is expected to run from 1 June 2018 to 20 September 2018. However, this schedule may change due to unanticipated circumstances, and thus the chart will be used as a guideline.
Impact
This project is expected to have a vast impact, including academic, personal, industrial, societal, and economic outcomes. The completion of this project will enhance the personal knowledge of the student and contribute to their research and career growth. This project will provide an opportunity for the student to learn and appreciate computational modelling. The findings of the project will be published in peer-reviewed journals, thus permitting the student to disseminate the findings of their work to other researchers in the field.
This work will shed more light on the versatility of the cytochrome P450 enzyme, which may inform future studies to exploit this group of enzymes in the production of other useful substances. This feat will bring personal satisfaction, a personal sense of achievement and contribution to the common good of society. The successful conclusion of this venture will provide academic benefits by enabling the master’s student to develop a detailed, high-quality dissertation. Consequently, the student will be able to fulfil the academic requirements of the University of Manchester for the award of a master’s degree.
Industries can benefit from this work by switching to cost-effective and sustainable energy sources for their processes. The findings of this study can be implemented on a large scale and commercialized to produce large quantities of biofuel. Therefore, new industries specializing in the production of biofuels can emerge.
The combustion of petroleum from fossil fuels has adverse environmental outcomes. This process emits significant amounts of carbon monoxide gas, which pollutes the environment, adds to greenhouse gases, and leads to global warming, among other adverse consequences. The productive implementation of this project will lead to the development of an environmentally friendly source of fuel and reduce the levels of pollution associated with the combustion of petroleum fuels.
This move will result in societal benefits by reducing the burden of air, soil, and water pollution. Pollution has negative effects on human and animal health, including respiratory complications (air pollution) and gastrointestinal problems (water and soil pollution). Biofuel use will eliminate these health concerns and lead to healthier populations. However, as much as minimising the emission of greenhouse gases is among the key motivations for the development of biofuels, other unanticipated effects in terms of soil attributes, biodiversity, and water quality may arise. Therefore, it may be necessary to conduct thorough assessments concerning the environmental impact of terminal alkene production by OleTJE.
Economic benefits include the ability to generate large quantities of biofuel. Previous methods of generating terminal olefins using NAD(P)H and hydrogen peroxide are not economically viable due to the prohibitive cost of reducing power and large-scale use of hydrogen peroxide. The successful completion of this project will enable the engineering of OleTJE to produce large quantities of biofuel through a peroxide-independent pathway, which will eliminate the need for hydrogen peroxide. Instead, molecular oxygen, which is readily available, will be used as the oxidant. Current methods of producing biofuel are still more expensive than the cost of producing fossil fuel.
Engineering OleTJE to overproduce biofuels using cost-effective methods will make the production of biofuels sustainable and affordable, which will encourage people to use them for their energy needs. Additionally, eliminating the undesired α- and β-hydroxylation products will produce large quantities of terminal olefins for use as biofuel. In this way, concerns about the escalating costs of crude oil and the increasing consumption of fuel will be addressed by the cost-efficient production of biofuels. Countries with little or no fossil fuel reserves will be able to enjoy energy security through the adoption of this technology.
The advancement of biofuels provides opportunities and challenges for developing countries. Non-oil producing nations may obtain partial substitutes for expensive oil imports. This venture may also provide an extra source of income and contribute towards rural development and enhancement of local infrastructures. Other possible economic benefits include the emergence of biofuel industries, which will lead to economic growth through job creation and the sale of raw materials (feedstock) and products. However, negative effects, such as the depletion of water resources, food insecurity, and pollution, may also occur. Therefore, detailed cost-benefit analyses are necessary before embarking on large-scale biofuel production ventures.
References
Belcher, J, McLean, KJ, Matthews, S, Woodward, LS, Fisher, K, Rigby, SE, Nelson, DR, Potts, D, Baynham, MT, Parker, DA & Leys, D 2014, ‘Structure and biochemical properties of the alkene producing cytochrome P450 OleTJE (CYP152L1) from the Jeotgalicoccus sp. 8456 bacterium’, Journal of Biological Chemistry, vol. 289, no. 10, pp. 6535-6550.
Chen, B, Lee, DY & Chang, MW 2015, ‘Combinatorial metabolic engineering of Saccharomyces cerevisiae for terminal alkene production’, Metabolic Engineering, vol. 31, pp. 53-61.
Covert, T, Greenstone, M & Knittel, CR 2016, ‘Will we ever stop using fossil fuels?’ Journal of Economic Perspectives, vol. 30, no. 1, pp. 117-138.
de Visser, SP 2009, ‘Introduction to quantum behaviour – a primer,’ in RK Allemann & NS Scrutton (eds), Quantum tunnelling in enzyme-catalysed reactions, Royal Society of Chemistry, Cambridge, UK, pp. 18-35.
de Visser, SP, Quesne, MG, Martin, B, Comba, P & Ryde, U 2014, ‘Computational modelling of oxygenation processes in enzymes and biomimetic model complexes’, Chemical Communications, vol. 50, no. 3, pp. 262-282.
Dennig, A, Kuhn, M, Tassoti, S, Thiessenhusen, A, Gilch, S, Bülter, T, Haas, T, Hall, M & Faber, K 2015, ‘Oxidative decarboxylation of short‐chain fatty acids to 1‐alkenes’, Angewandte Chemie International Edition, vol. 54, no. 30, pp. 8819-8822.
Duarte, F, Amrein, BA, Blaha-Nelson, D & Kamerlin, SC 2015, ‘Recent advances in QM/MM free energy calculations using reference potentials’, Biochimica et Biophysica Acta (BBA)-General Subjects, vol. 1850, no. 5, pp. 954-965.
Fang, B, Xu, H, Liu, Y, Qi, F, Zhang, W, Chen, H, Wang, C, Wang, Y, Yang, W & Li, S 2017, ‘Mutagenesis and redox partners analysis of the P450 fatty acid decarboxylase OleT JE’, Scientific Reports, vol. 7, p. 44258.
Faponle, AS, Quesne, MG & de Visser, SP 2016, ‘Origin of the regioselective fatty‐acid hydroxylation versus decarboxylation by a cytochrome P450 Peroxygenase: what drives the reaction to biofuel production?’ Chemistry-A European Journal, vol. 22, no. 16, pp. 5478-5483.
Grant, JL, Hsieh, CH & Makris, TM 2015, ‘Decarboxylation of fatty acids to terminal alkenes by cytochrome P450 compound I’, Journal of the American Chemical Society, vol. 137, no. 15, pp. 4940-4943.
Hayashi, S, Uchida, Y, Hasegawa, T, Higashi, M, Kosugi, T & Kamiya, M 2017, ‘QM/MM geometry optimization on extensive free-energy surfaces for examination of enzymatic reactions and design of novel functional properties of proteins’, Annual Review of Physical Chemistry, vol. 68, pp. 135-154.
Ji, L, Faponle, AS, Quesne, MG, Sainna, MA, Zhang, J, Franke, A, Kumar, D, van Eldik, R, Liu, W & de Visser, SP 2015, ‘Drug metabolism by cytochrome P450 enzymes: what distinguishes the pathways leading to substrate hydroxylation over desaturation?’ Chemistry-A European Journal, vol. 21, no. 25, pp. 9083-9092.
Jindal, G & Warshel, A 2016, ‘Exploring the dependence of QM/MM calculations of enzyme catalysis on the size of the QM region’, The Journal of Physical Chemistry B, vol. 120, no. 37, pp. 9913-9921.
Lewars, EG 2016, Computational chemistry: introduction to the theory and applications of molecular and quantum mechanics, Springer, New York.
Liao, JC, Mi, L, Pontrelli, S & Luo, S 2016, ‘Fuelling the future: microbial engineering for the production of sustainable biofuels’, Nature Reviews Microbiology, vol. 14, no. 5, pp. 288-304.
Lin, FM, Marsh, ENG & Lin, XN 2015, ‘Recent progress in hydrocarbon biofuel synthesis: pathways and enzymes’, Chinese Chemical Letters, vol. 26, no. 4, pp. 431-434.
Liu, Y, Wang, C, Yan, J, Zhang, W, Guan, W, Lu, X & Li, S 2014, ‘Hydrogen peroxide-independent production of α-alkenes by OleT JE P450 fatty acid decarboxylase’, Biotechnology for Biofuels, vol. 7, no. 1, p. 28.
Matthews, S, Belcher, JD, Tee, KL, Girvan, HM, McLean, KJ, Rigby, SE, Levy, CW, Leys, D, Parker, DA, Blankley, RT & Munro, AW 2017a, ‘Catalytic determinants of alkene production by the cytochrome P450 peroxygenase OleTJE’, Journal of Biological Chemistry, vol. 292, no. 12, pp. 5128-5143.
Matthews, S, Tee, KL, Rattray, NJ, McLean, KJ, Leys, D, Parker, DA, Blankley, RT & Munro, AW 2017b, ‘Production of alkenes and novel secondary products by P450 OleTJE using novel H2O2‐generating fusion protein systems’, FEBS Letters, vol. 591, no. 5, pp. 737-750.
Melander, M, Jónsson, EO, Mortensen, JJ, Vegge, T & García Lastra, JM 2016, ‘Implementation of constrained DFT for computing charge transfer rates within the projector augmented wave method’, Journal of Chemical Theory and Computation, vol. 12, no. 11, pp. 5367-5378.
Munro, AW, McLean, KJ, Grant, JL & Makris, TM 2018, ‘Structure and function of the cytochrome P450 peroxygenase enzymes’, Biochemical Society Transactions, vol. 46, no. 1, pp. 183-196.
Nicoletti, G, Arcuri, N, Nicoletti, G & Bruno, R 2015, ‘A technical and environmental comparison between hydrogen and some fossil fuels’, Energy Conversion and Management, vol. 89, pp. 205-213.
Quesne, MG, Borowski, T & de Visser, SP 2016, ‘Quantum mechanics/molecular mechanics modelling of enzymatic processes: caveats and breakthroughs,’ Chemistry–A European Journal, vol. 22, no. 8, pp. 2562-2581.
Wang, J & Zhu, K 2018, ‘Microbial production of alka (e) ne biofuels’, Current Opinion in Biotechnology, vol. 50, pp. 11-18.
Xu, H, Ning, L, Yang, W, Fang, B, Wang, C, Wang, Y, Xu, J, Collin, S, Laeuffer, F, Fourage, L & Li, S 2017, ‘In vitro oxidative decarboxylation of free fatty acids to terminal alkenes by two new P450 peroxygenases’, Biotechnology for Biofuels, vol. 10, no. 1, p. 208.
Zargar, A, Bailey, CB, Haushalter, RW, Eiben, CB, Katz, L & Keasling, JD 2017, ‘Leveraging microbial biosynthetic pathways for the generation of ‘drop-in’ biofuels’, Current Opinion in Biotechnology, vol. 45, pp. 156-163.
Zhao, YJ, Cheng, QQ, Su, P, Chen, X, Wang, XJ, Gao, W, & Huang, LQ 2014, ‘Research progress relating to the role of cytochrome P450 in the biosynthesis of terpenoids in medicinal plants’, Applied Microbiology and Biotechnology, vol. 98, no. 6, pp. 2371-2383.