- Introduction
- Tracing of fluorophore-labeled molecules (Aflatoxin M1 in milk)
- Primer extension reaction-based studies
- SPFS spectroscopy as applied in the interfacial hybridization analysis
- SPFS-SPRM as used in surface imaging
- The use of SPFS spectroscopy in analysis of antibody-antigen interaction
- SPFS as applied in colloids
- Conclusion
- References
Introduction
Surface plasmon resonance spectroscopy (SPR) represents an optical technique that is widely used in the classification of both thin films and interfaces. The underlying principles that are synonymous with this technique are widely reported in literature. Analyses that benefit from the sensitivity of this technique are those that are regarded to as bio-active in nature.
At this juncture, we have a reaction pair such that one partner chemically attaches itself to the biofunctional layer and the other partner carries the surface plasmon mode while maintaining contact with the analyte. As such, as the analyte molecules leave the solution they impart a corresponding change in the refractive index. This can be scrutinized in real-time and in an operation that is label-free. In a nutshell, a quantifiable sensor signal is generated once the analyte molecules are detected.
However, one setback of this technique is that the sensor signal might go unnoticed when the analyte enrichment (density) is extremely low or its molecules are tiny to generate signals. Among the techniques adopted to limit this problem is the application of a “quasi-three dimensional surface layer of a hydrogel or a polymer brush probed by surface plasmon wave leading to effective increase in binding site density” (Ford & Weber, 2013). Even so, many relevant analyte concentrations go undetected.
Recently, the launch of surface plasmon field-enhanced fluorescence spectroscopy (SPFS) has functioned to completely eliminate this detection limit.
The underlying principle adopted in this technique is that a superior electromagnetic field emanating from a surface plasmon is used in the excitation of surface-bound fluorophores (Wolfgang & Neumann, 2013). As a result, the emitted fluorescence is eventually analyzed to reflect the behavior of the analyte. The figure below is a schematic representation of the SPFS (Stewart & Thompsons, 2008).
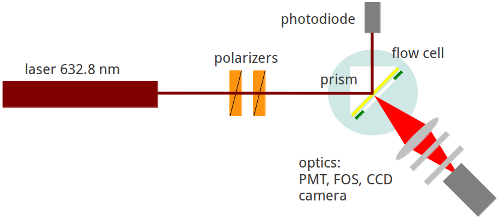
Since its inception, SPFS technique has been used to achieve a number of milestones. It has successfully been used to: trace fluorophore-labeled molecules, analyze primer extension reaction-based studies, study interfacial hybridization analysis, perform imaging of surfaces, study antibody-antigen interaction, and perform colloids studies. However, its importance is not limited to the aforementioned applications. There are still many more areas where this technique has excelled. For the sake of this review, we limit our scope to the above mentioned applications.
Tracing of fluorophore-labeled molecules (Aflatoxin M1 in milk)
Prior to the discovery of SPFS, the existing techniques were unable to probe miniature quantities of analytes in real-time. As a consequence, these techniques were rendered useless in analyses that required stringent measures, for instance, proteomics. Since its inception in the year 1999, SPFS has made possible these analyses giving accurate and reliable results.
Apart from the conventional SPR spectroscopy probing of the thickness changes at the interface, SPFS offers “the fluorescence signal channel that can directly monitor biomolecular binding kinetics in a highly sensitive way” (Duque et al., 2012). Nonetheless, the incident fluorescence that emanate from the flourophores experience considerable quenching by metals in the order of less than 10 nm dye-to-metal distances.
This limits the benefits that are achieved from the SPR field effects. Moreover, an imminent signal deviation, an undesirable effect is anticipated. This signal emanates from a combined effect of the fading “surface plasmon field and the metal-induced quenching” (Huang & Yu, 2013).
Recently, efforts have been put in place to reduce the problem posed by dye-to-metal distance. These strategies that include among others the layer-by-layer strategy are meant to optimize on the dye-to-metal allowance. For the sake of this analysis, a spatially extended matrix is adopted as a binding template vital in overcoming the aforementioned drawbacks.
In order to detect aflatoxin M1 in milk, a novel biosensor that is extremely sensitive is employed. The biosensor is basically based on an already advanced SPFS courtesy “of the excitation of long range surface plasmons (LRSPs)” (Fang & Bjorn, 2013). Whatever happens in the SPFS is that the flurophore-labeled elements get attached to the sensor surface. At the same time, this reaction is probed with surface plasmons (SPs) that result in fluorescence that sends signals to the detector.
Like it has been mentioned before, the principle adopted utilizes the enhanced field intensity that result from the excitation of surface plasmons to amplify the ensuing fluorescence signal. In order to achieve detection effect, both the LRSP-enhanced fluorescence spectroscopy and an inhibitor immunoassay ought to be merged.
At this juncture, the derivative aflatoxin M1 gets localized on the surface of the sensor. As such, the antibodies that are typically antagonistic to aflatoxin M1 come in handy as recognition elements. To this end, the aflatoxin M1 can easily be detected thanks to the biosensor.
Primer extension reaction-based studies
The enzyme DNA polymerase is one key catalyst that has been employed by biotechnologists in the in-vitro manufacture of double-stranded DNA. These enzymes come in handy in both DNA sequencing methodologies and in PCR-based procedures vital in pinpointing the occurrence of mutation.
Vitally, methodologies that employ unique enzyme functions to ascertain the “identity of a base sequence, such as DNA ligation, digestion, restriction and primer extension, are generally more sensitive to base aberrations than those based on DNA hybridization reactions alone” (Giannini et al., 2012). The current procedures are aimed at improving both the operational competence and the sensitivity limits. One of the commonly used approaches analyzes DNA microarrays by exploring the fluorescence intensities.
Contemporary procedures explore the surface attachment of the reacting compounds to execute their principles that are typically real-time methods. Among these procedures include quartz crystal microbalance (QCM) and surface plasmon resonance (SPR). These two have successfully been used to study both the RNA and DNA polymerase profiles.
Basically, this procedure is more versatile vis-à-vis the previous DNA microarrays method since it provides information on the enzyme kinetics. With this information one is better placed to understand the dynamics of the interaction. Even with their competences in analyzing DNA synthesis, they are short of monitoring nucleotide incorporation events. This is owed to the fact they fail to detect miniature corresponding mass changes.
SPFS methodology comes in handy in detecting these negligible mass changes. For the sake of this report, we review a study that brings to light how SPFS was used to examine the catalytic action involved in the integration of “fluorescently labeled nucleotides into surface-attached oligonucleotides” (Dostálek & Knoll, 2013).
Of note, the catalyst that is involved in this study is DNA polymerase I. In SPFS, a brief electromagnetic field emanating from the surface plasmon mode and transmitting parallel to the gold-water interface enhances the excitation of surface-bound flourophores. The resonance of excitation is dependent on the refractive index of the interfacial layer. This can be enhanced by altering the angle of incident of the excitation light.
At resonance, the resulting interfacial field is amplified two octaves higher than the incident light (when the interface λ is set at 633 nm) (Tawa & Knoll, 2013). The optimal strength that dies off exponentially to the surface is realized at the interface. The amplification of the evanescent field is what gives SPFS its great sensitivity, making it different from TIRF (total internal reflection fluorescence) spectroscopy.
SPFS spectroscopy as applied in the interfacial hybridization analysis
Advances made in the development of DNA biosensors that has enabled them exhibit unequalled sensitivity and selectivity is owed to the successes made in Human Genome Project (HGP). Some biosensors including acoustic waves, fiber optics and plasmon surface-based have successfully analyzed oligonucleotide, achieving immediate scrutiny of hybridization kinetics.
Nonetheless, these techniques have failed to merit in the detection of PCR-amplified DNA. Ideally, oligonucleotides and PCR products are dissimilar analytes differing chiefly in two aspects (Knoll, 2012). First, the difference comes in the respective lengths of the bases forming them. Basically, oligonucleotides analytes that are utilized in biosensing analysis are normally less than 30 bases in length. This is dwarfed by the length of the PCR products that accommodate several kilo bases (kb).
As such, this massive amount of bases coupled with complexity in sequencing has been a major hindrance in PCR product analysis. For example, some of the non-selective portions of the products may jeopardize the hybridization process by physically adhering to the surface-bound probes, exuding high background signals. Moreover, the existence of these non-selective pieces enhances the magnitude of steric hindrance which limits the effectiveness of binding of PCR products.
Second, typically, PCR products are double-stranded in nature. This is a major setback that bars the recognition units from accessing the surface-bound probes. As a result, the sensitivity of the analysis is greatly reduced. For these techniques, only the end-point results are reported. As such, these techniques do not offer an in-depth analysis of the hybridization mechanism. This can only be offered by SPFS (Robelek et al., 2013).
In this review, we analyze how SPFS can be used to inspect hybridization of PCR products using PNAs (peptide nucleic acids) probes. Here, the initial step was to clearly explain the importance of antisense strand as applied in hybridization. In this effect, three PCR products were developed and then distinguished by labeling. These included the sense, antisense and sense-antisense double strands.
These were then subjected to heat denaturation process prior to a hybridization process with surface-bound PNA (at 10 mM NaOH). These were then monitored by SPFS in real-time. The results obtained showed that the sense-antisense strands exhibited a slightly higher hybridization signal vis-à-vis the sense strand. The lower signal as portrayed by the sense strand was caused by the antisense strands that elicited weak fluorescence.
After denaturation process, the initially separated double strands re-associated courtesy of NaOH concentration (Kellis & Poulose, 2012). Principally, higher salt concentration inhibits repulsion between denatured strands.
As revealed by SPFS technique, only a few antisense strands were seen to have been bound on to the surface during the entire hybridization process. This gives a glimpse of the importance of SPFS technique as used in monitoring hybridization process. This process deals with decimal components that require the sensitivity that is provided for by the aforementioned technique.
SPFS-SPRM as used in surface imaging
Simply put, surface plasmons are characterized by charge-density oscillations that occur at the boundary adjoining to media. The resonance that happens at the surface plasmons is achieved when two wave-vectors (p-polarized light and that of the surface plasmons) match. This result in a decrease in photon flux emanating from the media boundary.
The fields generated “from the surface plasmons interact with the immediate media surrounding the interface, only to decay exponentially to the media, with the rate of decay depended on the media used” (Attridge et al., 2013). As it has clearly been explained by Attridge et al. (2013), SPRM (surface plasmon resonance microscopy) employs the activation of surface plasmons to concurrently inspect the near-superficial refractive index at several sites of a sample surface.
The generated field can also be employed to activate flurophores as exhibited by SPFS or SPRF. The merits of this technique is that it displays high degree of sensitivity in real-time, it amplifies the intensity of the incident field up to 80 times, and the reflected field comes in handy in suppressing unwarranted background light.
Previous SPRM/SPRF “quipped with lasers in their systems display excellent qualities including narrow bandwidth and enhanced power” (Huang & Yu, 2013). Nonetheless, the effectiveness of SPRM/SPRF in accomplishing imaging has greatly been hampered owing to speckle artifacts that are brought about by laser illumination. This decreases the ability of the double−system to concurrently track and identify specific regions.
For the sake of eliminating the aforementioned problems, the system assembly incorporate convectional laser pointers and acoustic transducer (anti-despeckle). Notably, dark Mylar is incorporated on the flow−cell to enhance coherence of the emitted excitation light.
This further goes on to enhance synchronization of measurement at specific locations on the flow−cell. This system is meant to “demonstrate a highly cost-effective system capable of simultaneously interrogating a sample under SPRM and SPRF, thus spatially discriminating between regions on the same sample” (Fort & Grésillon, 2013).
In a synopsis, with the above adjustments done on the double−system, the two can concurrently and successfully achieve imaging of surfaces. The entire experiment demonstrating how this is achieved is available in literature. There are available instruments in the market that can individually achieve some of the above effects. Nonetheless, the SPRM/SPRF system presents the user with an opportunity to experience all the effects in a single application that is highly cost-efficient.
The use of SPFS spectroscopy in analysis of antibody-antigen interaction
The underlying principle adopted by SPFS spectroscopy technique is that a superior electromagnetic field emanating from a surface plasmon is used to enhance the excitation of surface-bound fluorophores. Whatever gives SPFS technique an upper hand in its application in surface immunoreactions analyses is that it has the capacity to concurrently inspect both the interfacial thickness variations and the incandescence signals in real-time.
In view of this application, gold surfaces were activated courtesy of an admixture of self-assembled monolayer. This functioned to expose an antigen variably over a wide scale as regards its density. At this point, specific antibody-antigen reactions were registered for the anti-antigen antibody solutions traversing over the surfaces at high velocity thanks to SFPS-based immunoassays. Initial analysis investigated the significance of the “fluorophores Cy5 and Alexa Fluor 647 in SPFS-based immunoassays” (Cai & Jun, 2012).
The results revealed that Cy5 portrays an excellent self-quenching behavior, which negatively influence the quantitative measurements. Moreover, angular ‘detuning’ effects witnessed when taking the in−situ measurements as regards binding kinetics was confirmed to be a major hindrance. As such, they hamper with the incandescence signals at the point where huge SPR signals are recorded. In a synopsis, as it has been explained herein, SPFS-based immunoassays were vital in the analysis of the antibody-antigen interactions.
SPFS as applied in colloids
For this application the literature below describes the effect of merging surface plasmons and field-enhanced spectroscopy in sensor behavior. The resonance activation of PSP components at a metal/buffer boundary in flow−cell amplifies the optical field strength relative to the incident laser light. Fresnel formula confirmed that indeed for a gold-water boundary, the magnitude of the resulting field is enhanced 16 folds that of the incident light (Tawa & Morigaki, 2013).
The resulting field comes in handy in enhancing the sensitivity for inspecting binding effects of an analyte that traverses an aqueous phase at the functionalized gold-water interface to the detection sites (Liebermann & Knoll, 2012). This would happen on condition that the previously fluorescent-labeled analyte particles wonder within the locus of an exponentially decaying temporal field that is generated by the PSP mode.
This also bars these analyte molecules from approaching a metal, a recipe that is important in enhancing Forster quenching as regards the emitted fluorescence (Liebermann, 2012). In order to investigate the significance of SPFS, a quantitative analysis is done to investigate both the magnitude of fluorescence and the refractive index or layer thickness. The analysis answers the aforementioned questions by investigating the binding process of fluorescently-doped latex molecules (Sonnefraud, 2012).
These molecules possess extra superficial biotin-moieties that enable them to attach to a streptavidin phase at the gold-buffer boundary. The significance of SPFS technique is also echoed by its ability to detect fluorescence intensity emanating from negligible PSP resonance. To this end, this is exhibited by biotinylated chromophore which gives considerably dilute binding (Lakowicz et al., 2013).
Conclusion
In a conclusion, the achievements made by SPFS in optical techniques are unequalled. Since its inception, SPFS has taken the optical technique to new heights, making analyses that initially seemed difficult a success.
By simply exciting the surface-bound fluorophores courtesy of a superior electromagnetic field emanating from surface plasmons, the behavior of the analyte can be determined thanks to the emitted fluorescence. One of the merits that give SPFS an edge over other optical techniques is that it exhibits unprecedented sensitivity in real-time. To this effect, major milestones have been achieved and as such one can use this technique to investigate behaviors of tiny components at the molecular level and beyond.
References
Attridge, J., Daniels, P., Deacon, J., & Robinsons, D. (2012). “Sensitivity enhancement of optical immunosensors by the use of a surface-plasmon resonance fluoroimmunoassay.” Biosens. Bioelectron, 6 (3), 201–214.
Cai, W. & Jun, Y. (2012). “Plasmonics for extreme light concentration and manipulation.” Nat. Mater, 9 (3), 193–204.
Dostálek, J., & Knoll, W. (2013). “Biosensors based on surface plasmon-enhanced fluorescence spectroscopy.” Biointerphases, 3 (3), 12–22.
Duque, L., & Menges, B., Borros, S., & Förch, R. (2012). Biomacromolecules. Sydney, Australia: IDP Education Australia.
Fang, Y., & Bjorn, P.(2013). Attomolar Sensitivity in Bioassays Based on Surface Plasmon Fluorescence Spectroscopy. J. AM. CHEM. SOC., 9 (126), 29-30.
Ford, G., & Weber, W. (2013). “Electromagnetic interactions of molecules with metal surfaces.” Phys. Rep., 113 (4), 195–287.
Fort, E., & Grésillon, S. (2013). “Surface enhanced fluorescence.” J. Phys. D Appl. Phys., 41(1), 1-13.
Giannini, V., Fernández-Domínguez, A., Sonnefraud, Y., & Roschuk, R. (2012). “Controlling light localization and light-matter interactions with nanoplasmonics.” Small, 6 (22), 2498–2507.
Huang, B., & Yu, F. (2013). Surface plasmon resonance imaging using a high numerical aperture microscope objective. Anal. Chem., 79 (1), 2979–2983.
Kellis, T., & Poulose, J. (2012). Surface plasmon resonance/surface plasmon enhanced fluorescence: An optical technique for the detection of multicomponent macromolecular adsorption at the solid/liquid interface. Langmuir, 18 (1), 6319– 6323.
Knoll, W. (2012). Interfaces and thin films as seen by bound electromagnetic waves. Annual Review of Physical Chemistry, 49, 569–638.
Lakowicz, J., Chowdhury, M., Szmacinski, H., & Fu, Y. (2013) “Plasmoncontrolled fluorescence: a new paradigm in fluorescence spectroscopy.” Chem. Rev., 133 (10), 1308–1346.
Liebermann, T & Knoll, W. (2012). “Surface-plasmon field-enhanced fluorescence spectroscopy,” Colloids Surf. A Physicochem. Eng. Asp., 171 (3), 115–130.
Liebermann, T. (2012). Colloids and Surfaces A: Physicochem. Eng. Aspects 171, (1), 115–130.
Robelek, R., Niu, L., Schmid, E. L., & Knoll, W. (2013). Multiplexed hybridization detection of quantum dot-conjugated DNA sequences using surface plasmon enhanced fluorescence microscopy and spectrometry. Anal. Chem., 76, 6160– 6165.
Sonnefraud, Y. (2012). Surface-plasmon field-enhanced fluorescence spectroscopy. Colloids and Surfaces. Physicochemical and Engineering Aspects, 171, 115– 130.
Stewart, M., & Thompsons, B. (2013). “Nanostructured plasmonic sensors,” Chem. Rev., 108 (2), 494–521.
Tawa, K., & Knoll, W. (2013). Mismatching base-pair dependence of the kinetics of DNA-DNA hybridization studied by surface plasmon fluorescence spectroscopy. Nucleic Acids Research, 32, 2372–2377.
Tawa, K., & Morigaki, K. (2013). Substrate-supported phospholipid membranes studied by surface plasmon resonance and surface plasmon fluorescence spectroscopy. Biophys. J., 89 (1), 2750–2758.
Wolfgang, K., & Neumann, T. (2013). Principles and applications of surface plasmons field-enhanced spectroscopy. New York, NY: New York University Press.